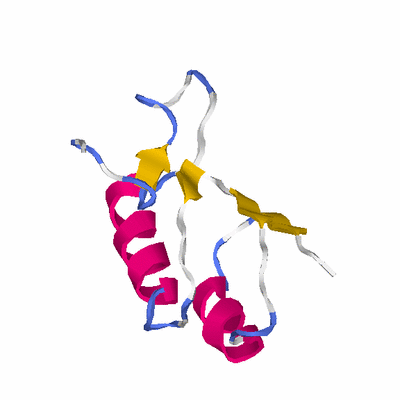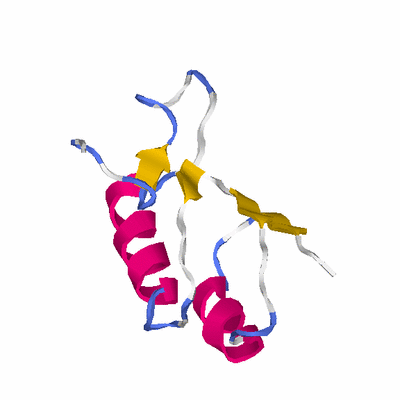##########
# PROTEIN: Q2246_545_101.5wLII_11068_74 126 101 # Category_C2
# Length: 126
# Weight: 14404.63
# Similarity: UniProt_B6AQP3, BLAST_B6AQP3, PDB_templates_INFO
# Best Pcover Model PDB N_AA SISC E-val Seq_ID LAL Overlap
# Cat 55.56 M_00 1o5z_A 442 1 0.026 30.000 70:75 (27-101:303-372)
# Evl 22.22 M_00 3bzx_B 83 1 0.014 35.000 28:28 (99-126:31-58)
# Sid 22.22 M_00 3bzx_B 83 1 0.014 35.000 28:28 (99-126:31-58)
# Cov 55.56 M_00 1o5z_A 442 1 0.026 30.000 70:75 (27-101:303-372)
# LaL 22.22 M_00 3bzx_B 83 1 0.014 35.000 28:28 (99-126:31-58)
# OvN 55.56 M_00 1o5z_A 442 1 0.026 30.000 70:75 (27-101:303-372)
# OvC 22.22 M_00 3bzx_B 83 1 0.014 35.000 28:28 (99-126:31-58)
PDB: 1o5z PDBsum
HEADER LIGASE 10-OCT-03 1O5Z
TITLE CRYSTAL STRUCTURE OF FOLYLPOLYGLUTAMATE SYNTHASE (TM0166)
TITLE 2 FROM THERMOTOGA MARITIMA AT 2.10 A RESOLUTION
SOURCE 2 ORGANISM_SCIENTIFIC: THERMOTOGA MARITIMA;
SOURCE 3 ORGANISM_TAXID: 2336;
KEYWDS TM0166, FOLYLPOLYGLUTAMATE SYNTHASE, STRUCTURAL GENOMICS,
KEYWDS 2 JCSG, PSI, PROTEIN STRUCTURE INITIATIVE, JOINT CENTER FOR
KEYWDS 3 STRUCTURAL GENOMICS, LIGASE
SCOP: c.59 MurD-like peptide ligases, peptide-binding domain
SCOP: c.59.1 MurD-like peptide ligases, peptide-binding domain
SCOP: c.59.1.2 Folylpolyglutamate synthetase, C-terminal domain
SCOP: c.72 Ribokinase-like
SCOP: c.72.2 MurD-like peptide ligases, catalytic domain
SCOP: c.72.2.2 Folylpolyglutamate synthetase
PDB: 3bzx PDBsum
HEADER MEMBRANE PROTEIN, PROTEIN TRANSPORT 18-JAN-08 3BZX
TITLE CRYSTAL STRUCTURE OF THE MUTATED H265A ESCU C-TERMINAL
TITLE 2 DOMAIN
SOURCE 2 ORGANISM_SCIENTIFIC: ESCHERICHIA COLI;
SOURCE 3 ORGANISM_TAXID: 562;
SOURCE 12 ORGANISM_SCIENTIFIC: ESCHERICHIA COLI;
KEYWDS AUTO CLEAVAGE PROTEIN, INTEIN, T3SS, MEMBRANE PROTEIN,
KEYWDS 2 PROTEIN TRANSPORT
SCOP: none