##########
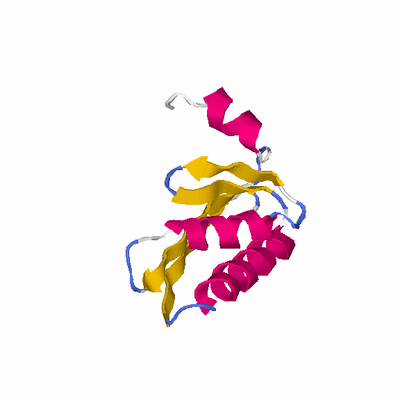
# PROTEIN: Q2246_545_226.5wLII_11195_17 104 226 # Category_A
# Length: 104
# Weight: 11702.46
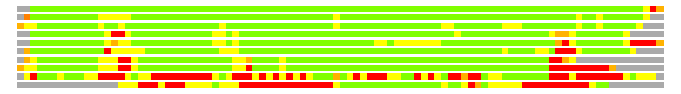
# Similarity: UniProt_A3ER18, BLAST_A3ER18, PDB_templates_INFO
# Best Pcover Model PDB N_AA SISC E-val Seq_ID LAL Overlap
# Cat 87.50 M_00 z0sd 104 0 2e-55 100.000 91:104 (1-104:1-104)
# Evl 87.50 M_00 z0sd 104 0 2e-55 100.000 91:104 (1-104:1-104)
# Sid 87.50 M_00 z0sd 104 0 2e-55 100.000 91:104 (1-104:1-104)
# Cov 95.19 M_02 1lxj_A 104 1 1e-21 24.000 99:101 (1-99:4-104)
# LaL 88.46 M_01 2epi_A 100 12 7e-24 36.000 92:92 (2-93:6-97)
# OvN 87.50 M_00 z0sd 104 0 2e-55 100.000 91:104 (1-104:1-104)
# OvC 87.50 M_00 z0sd 104 0 2e-55 100.000 91:104 (1-104:1-104)
PDB: 2epi PDBsum
HEADER STRUCTURAL GENOMICS, UNKNOWN FUNCTION 30-MAR-07 2EPI
TITLE CRYSTAL STRUCTURE PF HYPOTHETICAL PROTEIN MJ1052 FROM
TITLE 2 METHANOCALDOCOCCUS JANNASCII (FORM 2)
SOURCE 2 ORGANISM_SCIENTIFIC: METHANOCALDOCOCCUS JANNASCHII DSM
SOURCE 4 ORGANISM_TAXID: 243232;
KEYWDS NPPSFA, NATIONAL PROJECT ON PROTEIN STRUCTURAL AND
KEYWDS 2 FUNCTIONAL ANALYSES, RIKEN STRUCTURAL GENOMICS/PROTEOMICS
KEYWDS 3 INITIATIVE, RSGI, STRUCTURAL GENOMICS, UNKNOWN FUNCTION
SCOP: none
PDB: 1lxj PDBsum
HEADER STRUCTURAL GENOMICS, UNKNOWN FUNCTION 05-JUN-02 1LXJ
TITLE X-RAY STRUCTURE OF YBL001C NORTHEAST STRUCTURAL GENOMICS
TITLE 2 (NESG) CONSORTIUM TARGET YTYST72
SOURCE 2 ORGANISM_SCIENTIFIC: SACCHAROMYCES CEREVISIAE;
SOURCE 3 ORGANISM_COMMON: BAKER'S YEAST;
SOURCE 4 ORGANISM_TAXID: 4932;
KEYWDS HYPOTHETICAL PROTEIN, HTB2-NTH2 INTERGENIC REGION,
KEYWDS 2 STRUCTURAL GENOMICS, PSI, PROTEIN STRUCTURE INITIATIVE,
KEYWDS 3 NORTHEAST STRUCTURAL GENOMICS CONSORTIUM, NESG, UNKNOWN
KEYWDS 4 FUNCTION
SCOP: d.58 Ferredoxin-like
SCOP: d.58.48 MTH1187/YkoF-like
SCOP: d.58.48.1 MTH1187-like