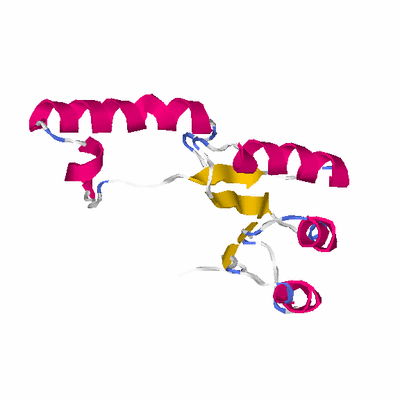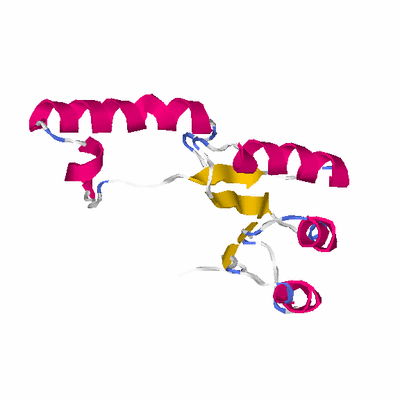##########
# PROTEIN: Q2246_545_324.5wLII_11276_28 173 324 # Category_C2
# Length: 173
# Weight: 18914.09
# Similarity: UniProt_B6AP32, BLAST_B6AP32, PDB_templates_INFO
# Best Pcover Model PDB N_AA SISC E-val Seq_ID LAL Overlap
# Cat 67.63 M_00 1wuf_A 393 2 0.007 10.000 117:128 (43-162:112-236)
# Evl 59.54 M_00 1wuf_A 393 2 0.001 13.000 103:122 (50-168:122-227)
# Sid 59.54 M_00 1wuf_A 393 2 0.001 13.000 103:122 (50-168:122-227)
# Cov 67.63 M_00 1wuf_A 393 2 0.007 10.000 117:128 (43-162:112-236)
# LaL 45.09 M_00 1d02_B 202 2 0.025 9.000 78:81 (94-171:21-101)
# OvN 67.63 M_00 1wuf_A 393 2 0.007 10.000 117:128 (43-162:112-236)
# OvC 45.09 M_00 1d02_B 202 2 0.025 9.000 78:81 (94-171:21-101)
PDB: 1wuf PDBsum
HEADER STRUCTURAL GENOMICS, UNKNOWN FUNCTION 07-DEC-04 1WUF
TITLE CRYSTAL STRUCTURE OF PROTEIN GI:16801725, MEMBER OF ENOLASE
TITLE 2 SUPERFAMILY FROM LISTERIA INNOCUA CLIP11262
SOURCE 2 ORGANISM_SCIENTIFIC: LISTERIA INNOCUA CLIP11262;
SOURCE 3 ORGANISM_TAXID: 272626;
KEYWDS STRUCTURAL GENOMICS, UNKNOWN FUNCTION, NYSGXRC TARGET T2186,
KEYWDS 2 ENOLASE SUPERFAMILY, PROTEIN STRUCTURE INITIATIVE, PSI, NEW
KEYWDS 3 YORK SGX RESEARCH CENTER FOR STRUCTURAL GENOMICS
SCOP: c.1 TIM beta/alpha-barrel
SCOP: c.1.11 Enolase C-terminal domain-like
SCOP: c.1.11.2 D-glucarate dehydratase-like
SCOP: d.54 Enolase N-terminal domain-like
SCOP: d.54.1 Enolase N-terminal domain-like
SCOP: d.54.1.1 Enolase N-terminal domain-like
PDB: 1d02 PDBsum
HEADER HYDROLASE/DNA 08-SEP-99 1D02
TITLE CRYSTAL STRUCTURE OF MUNI RESTRICTION ENDONUCLEASE IN
TITLE 2 COMPLEX WITH COGNATE DNA
SOURCE 7 ORGANISM_SCIENTIFIC: MYCOPLASMA;
SOURCE 8 ORGANISM_TAXID: 2093;
COMPND 9 EC: 3.1.21.4;
KEYWDS ALPHA/BETA PROTEIN, PROTEIN-DNA COMPLEX, DISTORTED DOUBLE
KEYWDS 2 HELIX, HYDROLASE/DNA COMPLEX
SCOP: c.52 Restriction endonuclease-like
SCOP: c.52.1 Restriction endonuclease-like
SCOP: c.52.1.8 Restriction endonuclease MunI