##########
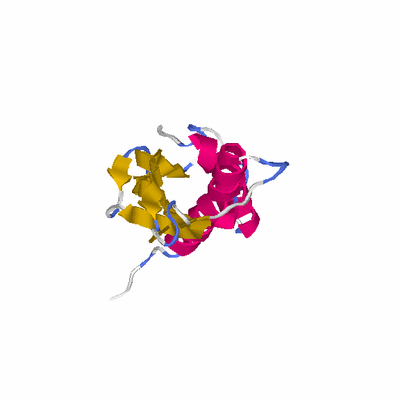
# PROTEIN: Q2246_545_367.5wLII_11277_49 196 367 # Category_C1
# Length: 196
# Weight: 20589.78
# Similarity: UniProt_B6ALQ7, BLAST_B6ALQ7, PDB_templates_INFO
# Best Pcover Model PDB N_AA SISC E-val Seq_ID LAL Overlap
# Cat 57.65 M_01 3bf2_A 152 1 2e-10 17.000 113:124 (33-156:25-142)
# Evl 68.88 M_00 2jxp_A 155 1 8e-25 12.000 135:147 (12-158:12-146)
# Sid 11.22 M_04 2i5n_M 323 14 0.098 50.000 22:22 (173-194:300-321)
# Cov 84.18 M_03 2z43_B 324 4 0.039 15.000 165:189 (6-173:37-222)
# LaL 11.22 M_04 2i5n_M 323 14 0.098 50.000 22:22 (173-194:300-321)
# OvN 71.43 M_00 2jxp_A 155 1 2e-19 11.000 140:154 (5-156:3-144)
# OvC 11.22 M_04 2i5n_M 323 14 0.098 50.000 22:22 (173-194:300-321)
PDB: 3bf2 PDBsum
HEADER LIPOPROTEIN 20-NOV-07 3BF2
TITLE CRYSTAL STRUCTURE OF THE A1KSW9_NEIMF PROTEIN FROM
TITLE 2 NEISSERIA MENINGITIDIS. NORTHEAST STRUCTURAL GENOMICS
TITLE 3 CONSORTIUM TARGET MR36A
SOURCE 2 ORGANISM_SCIENTIFIC: NEISSERIA MENINGITIDIS FAM18;
SOURCE 3 ORGANISM_TAXID: 272831;
KEYWDS LIPOPROTEIN, NESG, MR36A, STRUCTURAL GENOMICS, PSI-2,
KEYWDS 2 PROTEIN STRUCTURE INITIATIVE, NORTHEAST STRUCTURAL GENOMICS
KEYWDS 3 CONSORTIUM
SCOP: none
PDB: 2jxp PDBsum
HEADER LIPOPROTEIN 27-NOV-07 2JXP
TITLE SOLUTION NMR STRUCTURE OF UNCHARACTERIZED LIPOPROTEIN B
TITLE 2 FROM NITROSOMONAS EUROPAEA. NORTHEAST STRUCTURAL GENOMICS
TITLE 3 TARGET NER45A
SOURCE 2 ORGANISM_SCIENTIFIC: NITROSOMONAS EUROPAEA ATCC 19718;
SOURCE 3 ORGANISM_TAXID: 228410;
KEYWDS UNCHARACTERIZED LIPOPROTEIN B, NMR STRUCTURE, NORTHEAST
KEYWDS 2 STRUCTURAL GENOMICS CONSORTIUM, NESG, PSI-2, PROTEIN
KEYWDS 3 STRUCTURE INITIATIVE
SCOP: none
PDB: 2i5n PDBsum
HEADER PHOTOSYNTHESIS 25-AUG-06 2I5N
TITLE 1.96 A X-RAY STRUCTURE OF PHOTOSYNTHETIC REACTION CENTER
TITLE 2 FROM RHODOPSEUDOMONAS VIRIDIS:CRYSTALS GROWN BY
TITLE 3 MICROFLUIDIC TECHNIQUE
SOURCE 2 ORGANISM_SCIENTIFIC: BLASTOCHLORIS VIRIDIS;
SOURCE 3 ORGANISM_TAXID: 1079;
SOURCE 6 ORGANISM_SCIENTIFIC: BLASTOCHLORIS VIRIDIS;
KEYWDS PHOTOSYNTHETIC REACTION CENTER, SECONDARY QUINONE (QB),
KEYWDS 2 MICROFLUIDIC TECHNIQUE, HYBRID, MICROBATCH, PLUG,
KEYWDS 3 PHOTOSYNTHESIS
SCOP: a.138 Multiheme cytochromes
SCOP: a.138.1 Multiheme cytochromes
SCOP: a.138.1.2 Photosynthetic reaction centre (cytochrome subunit)
SCOP: b.41 PRC-barrel domain
SCOP: b.41.1 PRC-barrel domain
SCOP: b.41.1.1 Photosynthetic reaction centre, H-chain, cytoplasmic domain
SCOP: f.23 Single transmembrane helix
SCOP: f.23.10 Photosystem II reaction centre subunit H, transmembrane region
SCOP: f.23.10.1 Photosystem II reaction centre subunit H, transmembrane region
SCOP: f.26 Bacterial photosystem II reaction centre, L and M subunits
SCOP: f.26.1 Bacterial photosystem II reaction centre, L and M subunits
SCOP: f.26.1.1 Bacterial photosystem II reaction centre, L and M subunits
PDB: 2z43 PDBsum
HEADER RECOMBINATION 12-JUN-07 2Z43
TITLE STRUCTURE OF A TWINNED CRYSTAL OF RADA
SOURCE 2 ORGANISM_SCIENTIFIC: SULFOLOBUS SOLFATARICUS P2;
SOURCE 3 ORGANISM_TAXID: 273057;
KEYWDS ARCHAEA, FILAMENT, DNA BINDING, RECOMBINATION, MOLECULAR
KEYWDS 2 SWITCH, RECA, RAD51, DMC1
SCOP: none