##########
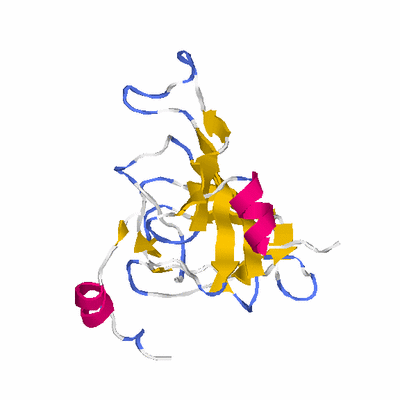
# PROTEIN: Q2246_545_524.5wLII_11390_10 173 524 # Category_B
# Length: 173
# Weight: 19612.32
# Similarity: UniProt_B6ARH9, BLAST_B6ARH9, PDB_templates_INFO
# Best Pcover Model PDB N_AA SISC E-val Seq_ID LAL Overlap
# Cat 83.82 M_00 1yud_A 170 10 2e-48 31.000 145:161 (13-171:12-158)
# Evl 83.82 M_00 1yud_A 170 10 2e-48 31.000 145:161 (13-171:12-158)
# Sid 71.68 M_00 1znp_A 154 7 3e-19 40.000 124:136 (14-145:6-133)
# Cov 91.91 M_03 1xe8_B 203 3 1e-27 23.000 159:176 (8-166:22-197)
# LaL 76.88 M_01 1yud_A 170 10 2e-46 36.000 133:139 (13-149:12-146)
# OvN 90.75 M_02 1xe7_B 203 3 1e-32 25.000 157:183 (1-166:23-197)
# OvC 83.82 M_00 1yud_A 170 10 2e-48 31.000 145:161 (13-171:12-158)
PDB: 1yud PDBsum
HEADER STRUCTURAL GENOMICS, UNKNOWN FUNCTION 14-FEB-05 1YUD
TITLE X-RAY CRYSTAL STRUCTURE OF PROTEIN SO0799 FROM SHEWANELLA
TITLE 2 ONEIDENSIS. NORTHEAST STRUCTURAL GENOMICS CONSORTIUM
TITLE 3 TARGET SOR12.
SOURCE 2 ORGANISM_SCIENTIFIC: SHEWANELLA ONEIDENSIS MR-1;
SOURCE 3 ORGANISM_TAXID: 211586;
KEYWDS X-RAY STRUCTURE, SOR12, Q8E1N8, PSI, PROTEIN STRUCTURE
KEYWDS 2 INITIATIVE, NORTHEAST STRUCTURAL GENOMICS CONSORTIUM, NESG,
KEYWDS 3 STRUCTURAL GENOMICS, UNKNOWN FUNCTION
SCOP: b.82 Double-stranded beta-helix
SCOP: b.82.1 RmlC-like cupins
SCOP: b.82.1.16 YML079-like
PDB: 1znp PDBsum
HEADER STRUCTURAL GENOMICS, UNKNOWN FUNCTION 11-MAY-05 1ZNP
TITLE X-RAY CRYSTAL STRUCTURE OF PROTEIN Q8U9W0 FROM
TITLE 2 AGROBACTERIUM TUMEFACIENS. NORTHEAST STRUCTURAL GENOMICS
TITLE 3 CONSORTIUM TARGET ATR55.
SOURCE 2 ORGANISM_SCIENTIFIC: AGROBACTERIUM TUMEFACIENS STR. C58;
SOURCE 3 ORGANISM_TAXID: 176299;
KEYWDS X-RAY, NESG, ATR55, Q8U9W0, STRUCTURAL GENOMICS, PSI,
KEYWDS 2 PROTEIN STRUCTURE INITIATIVE, NORTHEAST STRUCTURAL GENOMICS
KEYWDS 3 CONSORTIUM, UNKNOWN FUNCTION
SCOP: b.82 Double-stranded beta-helix
SCOP: b.82.1 RmlC-like cupins
SCOP: b.82.1.16 YML079-like
PDB: 1xe8 PDBsum
HEADER STRUCTURAL GENOMICS, UNKNOWN FUNCTION 09-SEP-04 1XE8
TITLE CRYSTAL STRUCTURE OF THE YML079W PROTEIN FROM SACCHAROMYCES
TITLE 2 CEREVISIAE REVEALS A NEW SEQUENCE FAMILY OF THE JELLY ROLL
TITLE 3 FOLD.
SOURCE 2 ORGANISM_SCIENTIFIC: SACCHAROMYCES CEREVISIAE;
SOURCE 3 ORGANISM_COMMON: BAKER'S YEAST;
SOURCE 4 ORGANISM_TAXID: 4932;
KEYWDS JELLY ROLL MOTIF, CUPIN SUPERFAMILY, STRUCTURAL GENOMICS,
KEYWDS 2 YML079WP, S. CEREVISIAE, UNKNOWN FUNCTION
SCOP: b.82 Double-stranded beta-helix
SCOP: b.82.1 RmlC-like cupins
SCOP: b.82.1.16 YML079-like
PDB: 1xe7 PDBsum
HEADER STRUCTURAL GENOMICS, UNKNOWN FUNCTION 09-SEP-04 1XE7
TITLE CRYSTAL STRUCTURE OF THE YML079W PROTEIN FROM SACCHAROMYCES
TITLE 2 CEREVISIAE REVEALS A NEW SEQUENCE FAMILY OF THE JELLY ROLL
TITLE 3 FOLD
SOURCE 2 ORGANISM_SCIENTIFIC: SACCHAROMYCES CEREVISIAE;
SOURCE 3 ORGANISM_COMMON: BAKER'S YEAST;
SOURCE 4 ORGANISM_TAXID: 4932;
KEYWDS JELLY ROLL MOTIF, CUPIN SUPERFAMILY, STRUCTURAL GENOMICS,
KEYWDS 2 YML079WP, S. CEREVISIAE, UNKNOWN FUNCTION
SCOP: b.82 Double-stranded beta-helix
SCOP: b.82.1 RmlC-like cupins
SCOP: b.82.1.16 YML079-like