LGA
Sequence Independent Analysis (LGA)
Frame of reference: Cat.Q2246_545_106.5wLII_11068_88
Total number of 3D structures: 20
LGA calculations using distance cutoff DIST: 4.0 A
Residues superimposed below 2.00 A: GREEN
Residues superimposed below 4.00 A: YELLOW
Residues superimposed below 6.00 A: ORANGE
Residues superimposed below 8.00 A: BROWN
Residues superimposed above 8.00 A or not aligned: RED
Terminal residues not aligned: GREY
Structure Deviation Summary
Calculations based on one final LGA superposition
(Bar representation of 3D plots, TEXT)
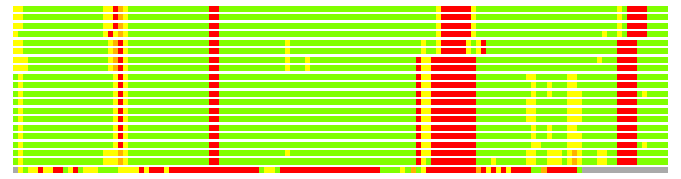
Structures ordered by LGA_S - score
| Structure |
NS |
NT |
N(dist=4.0) |
RMSD(N) |
Seq_ID(N) |
LGA_S |
LGA_Q |
PLOTS |
| 2vqm_A |
391 |
130 |
117 |
1.04 |
17.95 |
87.928 |
10.251 |
T P |
| 2vqo_A |
360 |
130 |
117 |
1.05 |
17.95 |
87.868 |
10.162 |
T P |
| 2vqq_A |
361 |
130 |
117 |
1.04 |
17.95 |
87.868 |
10.219 |
T P |
| 2vqw_G |
382 |
130 |
117 |
1.06 |
17.95 |
87.682 |
10.076 |
T P |
| 2nvr_A |
386 |
130 |
117 |
1.08 |
18.80 |
87.584 |
9.926 |
T P |
| 2pqp_A |
386 |
130 |
117 |
1.07 |
18.80 |
87.578 |
9.964 |
T P |
| 1zz1_A |
367 |
130 |
113 |
1.24 |
19.47 |
83.580 |
8.416 |
T P |
| 2vcg_D |
375 |
130 |
113 |
1.26 |
19.47 |
83.424 |
8.335 |
T P |
| 3ewf_A |
364 |
130 |
113 |
1.25 |
12.39 |
83.131 |
8.381 |
T P |
| 2v5w_B |
367 |
130 |
113 |
1.24 |
12.39 |
83.128 |
8.434 |
T P |
| 3ezp_A |
357 |
130 |
113 |
1.29 |
12.39 |
83.030 |
8.126 |
T P |
| 3f0r_B |
366 |
130 |
113 |
1.26 |
12.39 |
83.030 |
8.286 |
T P |
| 1t64_A |
364 |
130 |
113 |
1.26 |
12.39 |
82.955 |
8.333 |
T P |
| 2v5x_A |
363 |
130 |
113 |
1.28 |
12.39 |
82.777 |
8.162 |
T P |
| 3ew8_A |
357 |
130 |
113 |
1.28 |
12.39 |
82.726 |
8.186 |
T P |
| 3f06_A |
356 |
130 |
113 |
1.31 |
12.39 |
82.686 |
7.995 |
T P |
| 3ezt_A |
357 |
130 |
113 |
1.32 |
12.39 |
82.581 |
7.983 |
T P |
| 1c3r_A |
372 |
130 |
114 |
1.48 |
15.79 |
82.055 |
7.236 |
T P |
| 1c3p_A |
372 |
130 |
114 |
1.48 |
15.79 |
81.994 |
7.215 |
T P |
| 2jbr_A |
399 |
130 |
43 |
2.59 |
0.00 |
21.861 |
1.599 |
T P |
NS : Total number of residues in Structure (rotated structure)
NT : Total number of residues in TARGET (frame of reference)
N : Total number of residues superimposed under 4.0 Angstrom distance cutoff
RMSD : RMS deviation calculated on all N residues superimposed under 4.0 Angstrom distance cutoff
Seq_Id : Sequence Identity. Percent of identical residues from the total of N aligned.
LGA_S : Structure similarity score calculated by internal LGA procedure (see LGA paper for details)
LGA_Q : Score (how tight is the superposition) calculated by the formula: Q = 0.1*N/(0.1+RMSD)
PLOTS : T - Flat text file (output from LGA program, rotated structure)
PLOTS : P - Plot of superimposed structures (3D plot colored as bars)
Citing LGA:
Zemla A., "LGA - a Method for Finding 3D Similarities in Protein Structures",
Nucleic Acids Research, 2003, Vol. 31, No. 13, pp. 3370-3374.
[MEDLINE]

