LGA
Sequence Independent Analysis (LGA)
Frame of reference: Cat.Q2246_545_131.5wLII_11077_47
Total number of 3D structures: 99
LGA calculations using distance cutoff DIST: 4.0 A
Residues superimposed below 2.00 A: GREEN
Residues superimposed below 4.00 A: YELLOW
Residues superimposed below 6.00 A: ORANGE
Residues superimposed below 8.00 A: BROWN
Residues superimposed above 8.00 A or not aligned: RED
Terminal residues not aligned: GREY
Structure Deviation Summary
Calculations based on one final LGA superposition
(Bar representation of 3D plots, TEXT)
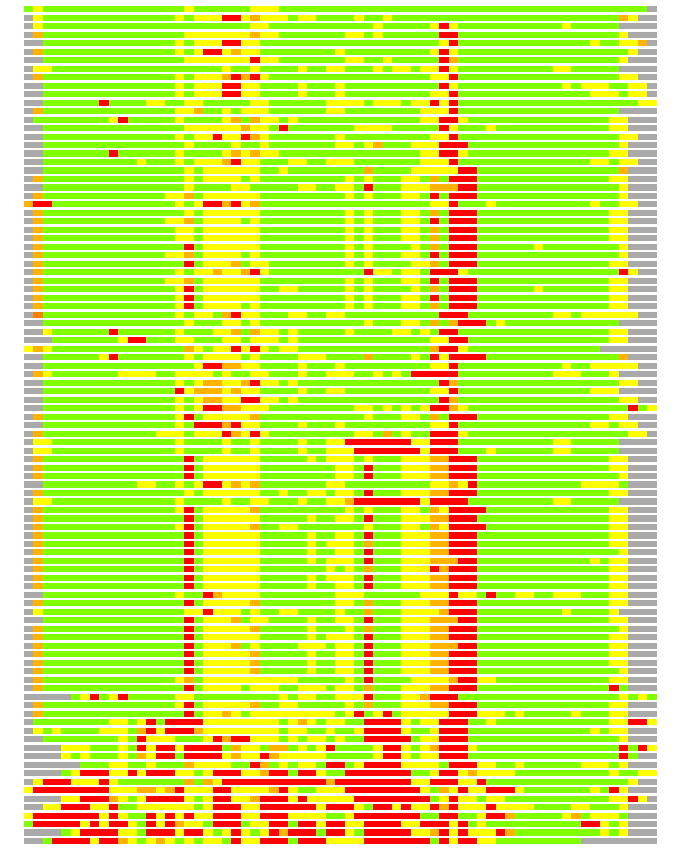
Structures ordered by LGA_S - score
| Structure |
NS |
NT |
N(dist=4.0) |
RMSD(N) |
Seq_ID(N) |
LGA_S |
LGA_Q |
PLOTS |
| 1wve_C |
75 |
67 |
66 |
0.96 |
31.82 |
96.498 |
6.240 |
T P |
| 3dp5_A |
99 |
67 |
62 |
1.69 |
20.97 |
85.847 |
3.456 |
T P |
| 1hzu_A |
521 |
67 |
61 |
1.59 |
22.95 |
85.042 |
3.607 |
T P |
| 1yiq_A |
684 |
67 |
61 |
1.85 |
22.95 |
84.508 |
3.132 |
T P |
| 2v08_A |
84 |
67 |
61 |
1.67 |
26.23 |
84.492 |
3.437 |
T P |
| 2zbo_A |
86 |
67 |
61 |
1.83 |
26.23 |
84.241 |
3.164 |
T P |
| 1kv9_A |
664 |
67 |
60 |
1.74 |
28.33 |
84.146 |
3.268 |
T P |
| 1e2r_B |
543 |
67 |
61 |
1.70 |
14.75 |
83.877 |
3.387 |
T P |
| 1gdv_A |
85 |
67 |
61 |
1.82 |
26.23 |
83.831 |
3.169 |
T P |
| 1kib_A |
89 |
67 |
61 |
1.73 |
21.31 |
83.458 |
3.331 |
T P |
| 1f1f_A |
88 |
67 |
61 |
1.76 |
21.31 |
83.379 |
3.283 |
T P |
| 1m70_A |
190 |
67 |
62 |
1.81 |
14.52 |
83.242 |
3.251 |
T P |
| 1nir_B |
539 |
67 |
61 |
1.78 |
21.31 |
82.495 |
3.253 |
T P |
| 1c52_A |
131 |
67 |
60 |
1.76 |
26.67 |
82.312 |
3.230 |
T P |
| 1kb0_A |
669 |
67 |
60 |
1.86 |
18.33 |
82.213 |
3.066 |
T P |
| 2d0w_A |
168 |
67 |
60 |
1.82 |
21.67 |
82.197 |
3.133 |
T P |
| 3cp5_A |
116 |
67 |
59 |
1.64 |
25.42 |
81.926 |
3.387 |
T P |
| 1dt1_A |
129 |
67 |
59 |
1.71 |
27.12 |
81.291 |
3.259 |
T P |
| 1c6r_A |
88 |
67 |
61 |
1.94 |
22.95 |
81.231 |
2.991 |
T P |
| 1j3s_A |
104 |
67 |
60 |
1.89 |
25.00 |
81.179 |
3.021 |
T P |
| 1cig_A |
108 |
67 |
60 |
1.92 |
21.67 |
81.140 |
2.973 |
T P |
| 1hro_A |
105 |
67 |
59 |
1.83 |
23.73 |
80.978 |
3.052 |
T P |
| 3cxh_W |
112 |
67 |
59 |
1.89 |
22.03 |
80.846 |
2.964 |
T P |
| 1ctj_A |
89 |
67 |
59 |
1.83 |
22.03 |
80.698 |
3.049 |
T P |
| 1cie_A |
108 |
67 |
60 |
1.93 |
21.67 |
80.636 |
2.961 |
T P |
| 1rap_A |
108 |
67 |
59 |
1.89 |
22.03 |
80.538 |
2.967 |
T P |
| 1cri_A |
108 |
67 |
60 |
1.96 |
21.67 |
80.532 |
2.906 |
T P |
| 1raq_A |
108 |
67 |
60 |
1.93 |
21.67 |
80.492 |
2.961 |
T P |
| 2b12_B |
108 |
67 |
59 |
1.92 |
22.03 |
80.073 |
2.916 |
T P |
| 1crh_A |
108 |
67 |
59 |
1.91 |
22.03 |
80.054 |
2.930 |
T P |
| 2aiu_A |
104 |
67 |
59 |
1.93 |
23.73 |
79.831 |
2.902 |
T P |
| 1umm_A |
149 |
67 |
58 |
1.95 |
24.14 |
79.633 |
2.830 |
T P |
| 1csx_A |
108 |
67 |
59 |
1.96 |
22.03 |
79.567 |
2.868 |
T P |
| 2b0z_B |
108 |
67 |
59 |
1.94 |
22.03 |
79.556 |
2.892 |
T P |
| 1s6v_B |
108 |
67 |
58 |
1.87 |
22.41 |
79.399 |
2.944 |
T P |
| 1ccr_A |
111 |
67 |
59 |
1.95 |
25.42 |
79.313 |
2.872 |
T P |
| 1ls9_A |
91 |
67 |
60 |
2.07 |
21.67 |
79.274 |
2.761 |
T P |
| 1fhb_A |
108 |
67 |
58 |
1.88 |
20.69 |
79.262 |
2.933 |
T P |
| 2fwl_A |
129 |
67 |
59 |
1.89 |
27.12 |
78.528 |
2.968 |
T P |
| 1foc_A |
128 |
67 |
57 |
1.77 |
26.32 |
77.459 |
3.049 |
T P |
| 1kx2_A |
81 |
67 |
57 |
1.88 |
17.54 |
77.410 |
2.882 |
T P |
| 1jdl_A |
118 |
67 |
56 |
1.78 |
30.36 |
77.258 |
2.984 |
T P |
| 2ce0_A |
99 |
67 |
60 |
1.91 |
26.67 |
75.292 |
2.983 |
T P |
| 1hzv_A |
514 |
67 |
57 |
1.97 |
21.05 |
74.655 |
2.757 |
T P |
| 1mg2_D |
147 |
67 |
61 |
2.01 |
18.03 |
72.743 |
2.887 |
T P |
| 2v07_A |
98 |
67 |
59 |
2.18 |
25.42 |
72.172 |
2.583 |
T P |
| 2gc4_D |
147 |
67 |
60 |
1.95 |
18.33 |
71.675 |
2.928 |
T P |
| 1w5c_T |
136 |
67 |
60 |
1.96 |
20.00 |
71.543 |
2.916 |
T P |
| 2b11_B |
108 |
67 |
59 |
2.00 |
22.03 |
71.452 |
2.803 |
T P |
| 1cyj_A |
90 |
67 |
58 |
1.77 |
25.86 |
71.365 |
3.101 |
T P |
| 1f1c_A |
129 |
67 |
60 |
1.93 |
20.00 |
71.090 |
2.962 |
T P |
| 1gq1_A |
559 |
67 |
52 |
1.81 |
17.31 |
70.948 |
2.729 |
T P |
| 1hj5_A |
559 |
67 |
52 |
1.84 |
17.31 |
70.542 |
2.682 |
T P |
| 1irw_A |
108 |
67 |
59 |
2.02 |
22.03 |
70.133 |
2.777 |
T P |
| 1ytc_A |
112 |
67 |
58 |
1.96 |
22.41 |
70.098 |
2.818 |
T P |
| 1cih_A |
108 |
67 |
58 |
1.98 |
22.41 |
70.009 |
2.785 |
T P |
| 2dge_A |
102 |
67 |
59 |
1.98 |
23.73 |
70.009 |
2.833 |
T P |
| 1irv_A |
108 |
67 |
59 |
2.03 |
22.03 |
69.989 |
2.773 |
T P |
| 1qks_A |
559 |
67 |
51 |
1.82 |
17.65 |
69.877 |
2.655 |
T P |
| 3cx5_W |
108 |
67 |
58 |
2.07 |
22.41 |
69.865 |
2.670 |
T P |
| 1yeb_A |
108 |
67 |
58 |
2.00 |
22.41 |
69.811 |
2.760 |
T P |
| 2bcn_B |
108 |
67 |
58 |
2.05 |
22.41 |
69.697 |
2.699 |
T P |
| 1yea_A |
112 |
67 |
58 |
2.00 |
22.41 |
69.687 |
2.763 |
T P |
| 1ctz_A |
108 |
67 |
59 |
2.04 |
22.03 |
69.601 |
2.763 |
T P |
| 1chi_A |
108 |
67 |
58 |
2.00 |
22.41 |
69.535 |
2.761 |
T P |
| 2ycc_A |
108 |
67 |
59 |
2.10 |
22.03 |
69.424 |
2.687 |
T P |
| 1csv_A |
108 |
67 |
58 |
2.00 |
20.69 |
69.411 |
2.763 |
T P |
| 1chj_A |
108 |
67 |
58 |
2.00 |
22.41 |
69.339 |
2.762 |
T P |
| 1csw_A |
108 |
67 |
58 |
2.01 |
22.41 |
69.313 |
2.752 |
T P |
| 1c6s_A |
87 |
67 |
59 |
2.03 |
23.73 |
69.219 |
2.766 |
T P |
| 1crg_A |
108 |
67 |
59 |
2.08 |
22.03 |
68.986 |
2.707 |
T P |
| 1yfc_A |
108 |
67 |
58 |
2.03 |
22.41 |
68.928 |
2.727 |
T P |
| 1wej_F |
104 |
67 |
58 |
1.98 |
24.14 |
68.782 |
2.786 |
T P |
| 1cif_A |
108 |
67 |
59 |
2.09 |
22.03 |
68.757 |
2.692 |
T P |
| 1chh_A |
108 |
67 |
58 |
2.05 |
22.41 |
68.698 |
2.701 |
T P |
| 2b4z_A |
104 |
67 |
59 |
2.09 |
23.73 |
68.684 |
2.693 |
T P |
| 1ycc_A |
108 |
67 |
58 |
2.04 |
22.41 |
68.586 |
2.708 |
T P |
| 2jqr_A |
108 |
67 |
58 |
2.04 |
22.41 |
68.586 |
2.708 |
T P |
| 1csu_A |
108 |
67 |
58 |
2.06 |
22.41 |
68.559 |
2.684 |
T P |
| 2jti_B |
103 |
67 |
60 |
2.20 |
21.67 |
68.537 |
2.614 |
T P |
| 1i54_A |
103 |
67 |
58 |
2.12 |
27.59 |
68.129 |
2.610 |
T P |
| 1h1o_A |
172 |
67 |
56 |
1.88 |
19.64 |
67.910 |
2.834 |
T P |
| 2b10_B |
108 |
67 |
59 |
2.15 |
22.03 |
64.110 |
2.625 |
T P |
| 1cyc_A |
103 |
67 |
57 |
2.23 |
29.82 |
60.881 |
2.444 |
T P |
| 1pby_A |
489 |
67 |
52 |
2.16 |
19.23 |
59.365 |
2.302 |
T P |
| 1jmx_A |
493 |
67 |
52 |
2.25 |
13.46 |
58.853 |
2.211 |
T P |
| 2blf_B |
81 |
67 |
50 |
1.98 |
8.00 |
58.432 |
2.404 |
T P |
| 2fw5_A |
125 |
67 |
48 |
2.17 |
12.50 |
55.027 |
2.113 |
T P |
| 2fwt_A |
112 |
67 |
48 |
2.26 |
8.33 |
54.016 |
2.034 |
T P |
| 1lms_A |
95 |
67 |
48 |
2.26 |
22.92 |
50.175 |
2.035 |
T P |
| 2j8s_A |
1044 |
67 |
40 |
2.38 |
2.50 |
42.926 |
1.614 |
T P |
| 2gif_B |
1044 |
67 |
37 |
2.36 |
2.70 |
42.693 |
1.501 |
T P |
| 2dhh_A |
1022 |
67 |
39 |
2.80 |
7.69 |
40.958 |
1.344 |
T P |
| 2hqd_A |
1016 |
67 |
39 |
2.56 |
0.00 |
40.607 |
1.467 |
T P |
| 2hqc_A |
1016 |
67 |
40 |
2.75 |
5.00 |
38.936 |
1.402 |
T P |
| 2hqg_A |
1016 |
67 |
36 |
2.54 |
5.56 |
38.608 |
1.363 |
T P |
| 2i6w_A |
1019 |
67 |
35 |
2.50 |
2.86 |
38.468 |
1.348 |
T P |
| 1t9x_A |
1014 |
67 |
36 |
2.52 |
0.00 |
37.294 |
1.372 |
T P |
| 2hqf_A |
1016 |
67 |
34 |
2.49 |
2.94 |
36.304 |
1.312 |
T P |
NS : Total number of residues in Structure (rotated structure)
NT : Total number of residues in TARGET (frame of reference)
N : Total number of residues superimposed under 4.0 Angstrom distance cutoff
RMSD : RMS deviation calculated on all N residues superimposed under 4.0 Angstrom distance cutoff
Seq_Id : Sequence Identity. Percent of identical residues from the total of N aligned.
LGA_S : Structure similarity score calculated by internal LGA procedure (see LGA paper for details)
LGA_Q : Score (how tight is the superposition) calculated by the formula: Q = 0.1*N/(0.1+RMSD)
PLOTS : T - Flat text file (output from LGA program, rotated structure)
PLOTS : P - Plot of superimposed structures (3D plot colored as bars)
Citing LGA:
Zemla A., "LGA - a Method for Finding 3D Similarities in Protein Structures",
Nucleic Acids Research, 2003, Vol. 31, No. 13, pp. 3370-3374.
[MEDLINE]

