LGA
Sequence Independent Analysis (LGA)
Frame of reference: Cat.Q2246_545_193.5wLII_11181_81
Total number of 3D structures: 43
LGA calculations using distance cutoff DIST: 4.0 A
Residues superimposed below 2.00 A: GREEN
Residues superimposed below 4.00 A: YELLOW
Residues superimposed below 6.00 A: ORANGE
Residues superimposed below 8.00 A: BROWN
Residues superimposed above 8.00 A or not aligned: RED
Terminal residues not aligned: GREY
Structure Deviation Summary
Calculations based on one final LGA superposition
(Bar representation of 3D plots, TEXT)
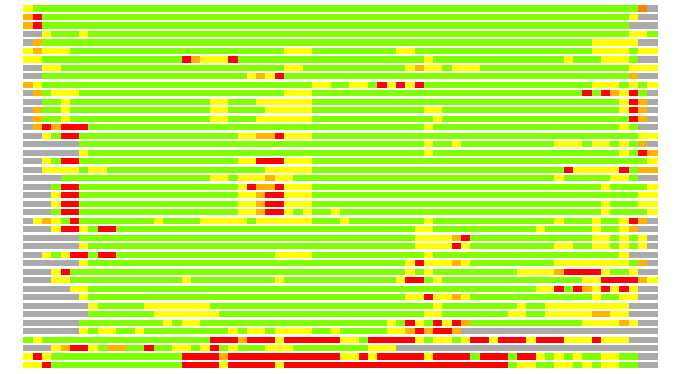
Structures ordered by LGA_S - score
| Structure |
NS |
NT |
N(dist=4.0) |
RMSD(N) |
Seq_ID(N) |
LGA_S |
LGA_Q |
PLOTS |
| 3f52_E |
78 |
68 |
67 |
1.23 |
17.91 |
96.214 |
5.055 |
T P |
| 2b5a_A |
77 |
68 |
65 |
0.90 |
23.08 |
95.698 |
6.497 |
T P |
| 1y7y_A |
69 |
68 |
64 |
0.72 |
25.00 |
94.324 |
7.805 |
T P |
| 1y9q_A |
178 |
68 |
66 |
1.11 |
19.70 |
94.163 |
5.444 |
T P |
| 3clc_B |
77 |
68 |
65 |
1.36 |
23.08 |
92.299 |
4.453 |
T P |
| 3f6w_A |
82 |
68 |
68 |
1.71 |
11.76 |
91.447 |
3.764 |
T P |
| 2ofy_A |
82 |
68 |
64 |
1.54 |
18.75 |
90.390 |
3.912 |
T P |
| 2bnm_A |
194 |
68 |
66 |
1.70 |
15.15 |
89.945 |
3.672 |
T P |
| 2ewt_A |
71 |
68 |
63 |
1.32 |
12.70 |
89.899 |
4.424 |
T P |
| 3b7h_A |
76 |
68 |
65 |
1.71 |
18.46 |
89.390 |
3.585 |
T P |
| 3eus_A |
85 |
68 |
63 |
1.52 |
14.29 |
88.829 |
3.882 |
T P |
| 1lmb_4 |
92 |
68 |
64 |
1.57 |
15.62 |
88.558 |
3.826 |
T P |
| 1rio_A |
97 |
68 |
65 |
1.72 |
16.92 |
88.513 |
3.565 |
T P |
| 1lli_B |
92 |
68 |
65 |
1.78 |
16.92 |
88.338 |
3.458 |
T P |
| 1adr_A |
76 |
68 |
61 |
1.35 |
14.75 |
87.371 |
4.193 |
T P |
| 2axz_A |
306 |
68 |
63 |
1.59 |
11.11 |
87.023 |
3.721 |
T P |
| 2r1j_L |
66 |
68 |
61 |
1.40 |
14.75 |
86.958 |
4.060 |
T P |
| 1b0n_A |
103 |
68 |
61 |
1.31 |
18.03 |
86.939 |
4.331 |
T P |
| 2aw6_A |
287 |
68 |
61 |
1.44 |
11.48 |
86.565 |
3.968 |
T P |
| 2ef8_A |
84 |
68 |
64 |
1.93 |
12.50 |
86.480 |
3.145 |
T P |
| 2qfc_A |
284 |
68 |
62 |
1.59 |
14.52 |
86.171 |
3.667 |
T P |
| 2grm_A |
315 |
68 |
61 |
1.54 |
11.48 |
85.771 |
3.721 |
T P |
| 2axu_A |
303 |
68 |
61 |
1.50 |
11.48 |
85.700 |
3.807 |
T P |
| 2awi_A |
298 |
68 |
61 |
1.51 |
11.48 |
85.410 |
3.783 |
T P |
| 2axv_B |
303 |
68 |
61 |
1.47 |
11.48 |
85.017 |
3.882 |
T P |
| 3bdn_A |
234 |
68 |
64 |
2.02 |
14.06 |
84.400 |
3.018 |
T P |
| 1utx_A |
66 |
68 |
59 |
1.46 |
22.03 |
83.778 |
3.775 |
T P |
| 2cro_A |
65 |
68 |
60 |
1.45 |
16.67 |
83.682 |
3.863 |
T P |
| 1r69_A |
63 |
68 |
60 |
1.52 |
15.00 |
82.430 |
3.711 |
T P |
| 3bs3_A |
62 |
68 |
59 |
1.48 |
22.03 |
82.018 |
3.735 |
T P |
| 1r63_A |
63 |
68 |
60 |
1.70 |
15.00 |
81.521 |
3.337 |
T P |
| 2jvl_A |
107 |
68 |
58 |
1.57 |
12.07 |
81.411 |
3.471 |
T P |
| 3dnv_B |
71 |
68 |
59 |
1.66 |
22.03 |
81.405 |
3.351 |
T P |
| 1x57_A |
91 |
68 |
57 |
1.35 |
15.79 |
80.676 |
3.920 |
T P |
| 1sq8_A |
64 |
68 |
59 |
1.71 |
11.86 |
80.650 |
3.267 |
T P |
| 2eby_A |
102 |
68 |
58 |
1.82 |
8.62 |
78.741 |
3.028 |
T P |
| 3cec_A |
91 |
68 |
58 |
1.92 |
15.52 |
78.477 |
2.876 |
T P |
| 2r63_A |
63 |
68 |
57 |
1.77 |
14.04 |
77.877 |
3.051 |
T P |
| 2k9q_A |
77 |
68 |
38 |
2.23 |
23.68 |
48.805 |
1.631 |
T P |
| 1jal_A |
348 |
68 |
39 |
2.12 |
7.69 |
45.140 |
1.758 |
T P |
| 3e7l_D |
63 |
68 |
33 |
2.37 |
15.15 |
41.279 |
1.337 |
T P |
| 3d5t_C |
323 |
68 |
34 |
1.99 |
11.76 |
40.122 |
1.628 |
T P |
| 1b8p_A |
327 |
68 |
32 |
1.90 |
12.50 |
37.383 |
1.599 |
T P |
NS : Total number of residues in Structure (rotated structure)
NT : Total number of residues in TARGET (frame of reference)
N : Total number of residues superimposed under 4.0 Angstrom distance cutoff
RMSD : RMS deviation calculated on all N residues superimposed under 4.0 Angstrom distance cutoff
Seq_Id : Sequence Identity. Percent of identical residues from the total of N aligned.
LGA_S : Structure similarity score calculated by internal LGA procedure (see LGA paper for details)
LGA_Q : Score (how tight is the superposition) calculated by the formula: Q = 0.1*N/(0.1+RMSD)
PLOTS : T - Flat text file (output from LGA program, rotated structure)
PLOTS : P - Plot of superimposed structures (3D plot colored as bars)
Citing LGA:
Zemla A., "LGA - a Method for Finding 3D Similarities in Protein Structures",
Nucleic Acids Research, 2003, Vol. 31, No. 13, pp. 3370-3374.
[MEDLINE]

