LGA
Sequence Independent Analysis (LGA)
Frame of reference: Cat.Q2246_545_195.5wLII_11184_6
Total number of 3D structures: 41
LGA calculations using distance cutoff DIST: 4.0 A
Residues superimposed below 2.00 A: GREEN
Residues superimposed below 4.00 A: YELLOW
Residues superimposed below 6.00 A: ORANGE
Residues superimposed below 8.00 A: BROWN
Residues superimposed above 8.00 A or not aligned: RED
Terminal residues not aligned: GREY
Structure Deviation Summary
Calculations based on one final LGA superposition
(Bar representation of 3D plots, TEXT)
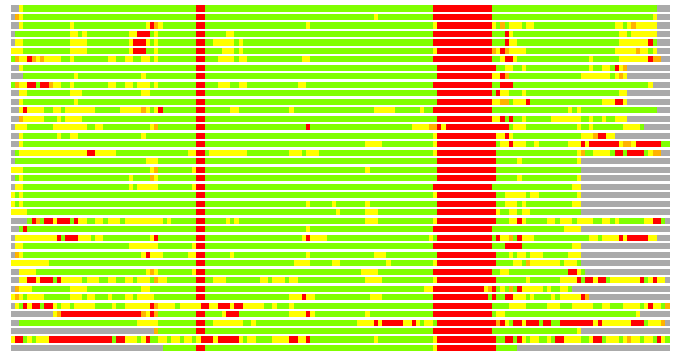
Structures ordered by LGA_S - score
| Structure |
NS |
NT |
N(dist=4.0) |
RMSD(N) |
Seq_ID(N) |
LGA_S |
LGA_Q |
PLOTS |
| 2fo7_A |
136 |
156 |
135 |
0.41 |
20.00 |
86.389 |
26.508 |
T P |
| 1w3b_A |
388 |
156 |
136 |
1.06 |
14.71 |
85.141 |
11.721 |
T P |
| 2vq2_A |
220 |
156 |
135 |
1.61 |
11.11 |
80.457 |
7.909 |
T P |
| 1e96_B |
185 |
156 |
132 |
1.40 |
7.58 |
80.038 |
8.793 |
T P |
| 1hh8_A |
192 |
156 |
131 |
1.41 |
6.87 |
79.127 |
8.653 |
T P |
| 1wm5_A |
205 |
156 |
134 |
1.65 |
8.21 |
79.080 |
7.640 |
T P |
| 2ho1_A |
222 |
156 |
134 |
1.91 |
8.21 |
78.599 |
6.668 |
T P |
| 3cv0_A |
300 |
156 |
127 |
1.25 |
13.39 |
78.379 |
9.373 |
T P |
| 2c0m_C |
302 |
156 |
127 |
1.45 |
14.96 |
78.065 |
8.195 |
T P |
| 2fi7_A |
223 |
156 |
129 |
1.67 |
7.75 |
77.516 |
7.299 |
T P |
| 2c0l_A |
292 |
156 |
128 |
1.33 |
14.84 |
77.513 |
8.953 |
T P |
| 3cvq_A |
289 |
156 |
126 |
1.33 |
14.29 |
77.095 |
8.805 |
T P |
| 2q7f_A |
194 |
156 |
133 |
1.81 |
14.29 |
77.047 |
6.950 |
T P |
| 1fch_A |
302 |
156 |
127 |
1.56 |
14.96 |
75.638 |
7.636 |
T P |
| 3edt_B |
258 |
156 |
130 |
1.77 |
11.54 |
75.523 |
6.965 |
T P |
| 2j9q_A |
300 |
156 |
122 |
1.43 |
15.57 |
75.056 |
7.971 |
T P |
| 2c2l_A |
281 |
156 |
123 |
1.63 |
8.13 |
74.584 |
7.120 |
T P |
| 1xnf_B |
262 |
156 |
129 |
2.02 |
7.75 |
73.667 |
6.094 |
T P |
| 1qz2_A |
285 |
156 |
118 |
1.07 |
9.32 |
73.650 |
10.063 |
T P |
| 1ihg_A |
364 |
156 |
119 |
1.21 |
9.24 |
73.434 |
9.064 |
T P |
| 1p5q_A |
283 |
156 |
118 |
1.10 |
9.32 |
73.251 |
9.852 |
T P |
| 1kt1_A |
374 |
156 |
118 |
1.14 |
9.32 |
73.043 |
9.548 |
T P |
| 2vyi_A |
128 |
156 |
119 |
1.29 |
14.29 |
72.495 |
8.561 |
T P |
| 1wao_1 |
471 |
156 |
119 |
1.35 |
8.40 |
72.177 |
8.185 |
T P |
| 1a17_A |
159 |
156 |
119 |
1.35 |
8.40 |
71.882 |
8.182 |
T P |
| 2pl2_A |
194 |
156 |
125 |
1.91 |
14.40 |
71.835 |
6.217 |
T P |
| 1na0_A |
119 |
156 |
116 |
1.16 |
18.10 |
71.798 |
9.200 |
T P |
| 3ceq_B |
269 |
156 |
123 |
1.83 |
13.01 |
71.261 |
6.362 |
T P |
| 1kt0_A |
357 |
156 |
114 |
1.21 |
10.53 |
70.251 |
8.675 |
T P |
| 2fbn_A |
153 |
156 |
117 |
1.54 |
6.84 |
69.813 |
7.140 |
T P |
| 2bug_A |
131 |
156 |
119 |
1.76 |
6.72 |
68.570 |
6.407 |
T P |
| 2dba_A |
148 |
156 |
116 |
1.60 |
10.34 |
68.410 |
6.824 |
T P |
| 2vsy_A |
547 |
156 |
124 |
2.07 |
11.29 |
68.404 |
5.714 |
T P |
| 1elw_A |
117 |
156 |
116 |
1.67 |
9.48 |
68.356 |
6.535 |
T P |
| 1elr_A |
128 |
156 |
117 |
1.91 |
8.55 |
68.258 |
5.835 |
T P |
| 2gw1_A |
487 |
156 |
127 |
2.12 |
11.02 |
64.787 |
5.713 |
T P |
| 2if4_A |
258 |
156 |
99 |
1.62 |
15.15 |
60.086 |
5.739 |
T P |
| 2vsn_A |
534 |
156 |
110 |
1.90 |
10.91 |
57.929 |
5.512 |
T P |
| 1na3_A |
86 |
156 |
85 |
1.05 |
21.18 |
53.512 |
7.406 |
T P |
| 1ouv_A |
265 |
156 |
103 |
2.25 |
13.59 |
50.229 |
4.387 |
T P |
| 2avp_A |
68 |
156 |
68 |
0.53 |
19.12 |
43.491 |
10.783 |
T P |
NS : Total number of residues in Structure (rotated structure)
NT : Total number of residues in TARGET (frame of reference)
N : Total number of residues superimposed under 4.0 Angstrom distance cutoff
RMSD : RMS deviation calculated on all N residues superimposed under 4.0 Angstrom distance cutoff
Seq_Id : Sequence Identity. Percent of identical residues from the total of N aligned.
LGA_S : Structure similarity score calculated by internal LGA procedure (see LGA paper for details)
LGA_Q : Score (how tight is the superposition) calculated by the formula: Q = 0.1*N/(0.1+RMSD)
PLOTS : T - Flat text file (output from LGA program, rotated structure)
PLOTS : P - Plot of superimposed structures (3D plot colored as bars)
Citing LGA:
Zemla A., "LGA - a Method for Finding 3D Similarities in Protein Structures",
Nucleic Acids Research, 2003, Vol. 31, No. 13, pp. 3370-3374.
[MEDLINE]

