LGA
Sequence Independent Analysis (LGA)
Frame of reference: Cat.Q2246_545_220.5wLII_11184_74
Total number of 3D structures: 30
LGA calculations using distance cutoff DIST: 4.0 A
Residues superimposed below 2.00 A: GREEN
Residues superimposed below 4.00 A: YELLOW
Residues superimposed below 6.00 A: ORANGE
Residues superimposed below 8.00 A: BROWN
Residues superimposed above 8.00 A or not aligned: RED
Terminal residues not aligned: GREY
Structure Deviation Summary
Calculations based on one final LGA superposition
(Bar representation of 3D plots, TEXT)
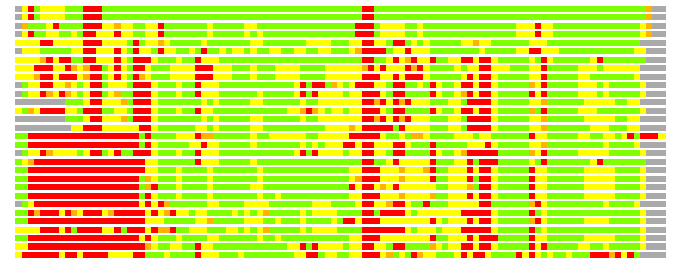
Structures ordered by LGA_S - score
| Structure |
NS |
NT |
N(dist=4.0) |
RMSD(N) |
Seq_ID(N) |
LGA_S |
LGA_Q |
PLOTS |
| 2cn3_A |
728 |
105 |
96 |
0.96 |
26.04 |
90.656 |
9.083 |
T P |
| 2cn2_A |
704 |
105 |
96 |
0.97 |
26.04 |
90.593 |
8.955 |
T P |
| 2ebs_B |
771 |
105 |
93 |
1.94 |
21.51 |
81.611 |
4.569 |
T P |
| 1sqj_B |
771 |
105 |
93 |
1.84 |
21.51 |
78.461 |
4.786 |
T P |
| 3b7f_A |
368 |
105 |
91 |
2.13 |
15.38 |
62.998 |
4.089 |
T P |
| 3f6k_A |
657 |
105 |
88 |
2.14 |
13.64 |
61.988 |
3.933 |
T P |
| 1dim_A |
381 |
105 |
81 |
2.00 |
14.81 |
60.665 |
3.851 |
T P |
| 1kit_A |
757 |
105 |
82 |
2.45 |
21.95 |
57.270 |
3.218 |
T P |
| 1w0p_A |
753 |
105 |
78 |
2.36 |
23.08 |
56.129 |
3.176 |
T P |
| 1s0i_A |
623 |
105 |
79 |
2.16 |
12.66 |
55.611 |
3.494 |
T P |
| 1ms5_A |
623 |
105 |
80 |
2.24 |
12.50 |
55.253 |
3.421 |
T P |
| 2vk5_A |
448 |
105 |
79 |
2.27 |
12.66 |
55.157 |
3.333 |
T P |
| 1ms9_B |
623 |
105 |
79 |
2.14 |
10.13 |
55.063 |
3.519 |
T P |
| 2bf6_A |
448 |
105 |
79 |
2.27 |
12.66 |
55.035 |
3.329 |
T P |
| 2w20_A |
470 |
105 |
77 |
2.09 |
12.99 |
54.726 |
3.520 |
T P |
| 2ber_A |
601 |
105 |
75 |
2.24 |
16.00 |
54.579 |
3.203 |
T P |
| 2jkb_A |
661 |
105 |
73 |
2.04 |
13.70 |
54.419 |
3.414 |
T P |
| 1mz5_A |
622 |
105 |
80 |
2.25 |
15.00 |
54.314 |
3.404 |
T P |
| 3sil_A |
379 |
105 |
73 |
1.90 |
15.07 |
54.045 |
3.656 |
T P |
| 1eut_A |
601 |
105 |
75 |
2.10 |
18.67 |
54.012 |
3.411 |
T P |
| 1eur_A |
361 |
105 |
76 |
2.17 |
19.74 |
53.896 |
3.345 |
T P |
| 1wcq_A |
601 |
105 |
76 |
2.20 |
18.42 |
53.819 |
3.305 |
T P |
| 1w8o_A |
601 |
105 |
76 |
2.17 |
18.42 |
53.494 |
3.341 |
T P |
| 2sli_A |
679 |
105 |
73 |
2.03 |
13.70 |
52.815 |
3.431 |
T P |
| 2ags_A |
634 |
105 |
75 |
2.33 |
20.00 |
52.394 |
3.083 |
T P |
| 2vvz_A |
470 |
105 |
73 |
2.22 |
19.18 |
52.190 |
3.148 |
T P |
| 1n1t_A |
628 |
105 |
75 |
2.30 |
21.33 |
51.584 |
3.119 |
T P |
| 2bq9_A |
601 |
105 |
74 |
2.15 |
18.92 |
50.827 |
3.291 |
T P |
| 1wcs_A |
628 |
105 |
70 |
2.08 |
12.86 |
50.387 |
3.216 |
T P |
| 2vw2_A |
658 |
105 |
70 |
2.28 |
7.14 |
48.185 |
2.943 |
T P |
NS : Total number of residues in Structure (rotated structure)
NT : Total number of residues in TARGET (frame of reference)
N : Total number of residues superimposed under 4.0 Angstrom distance cutoff
RMSD : RMS deviation calculated on all N residues superimposed under 4.0 Angstrom distance cutoff
Seq_Id : Sequence Identity. Percent of identical residues from the total of N aligned.
LGA_S : Structure similarity score calculated by internal LGA procedure (see LGA paper for details)
LGA_Q : Score (how tight is the superposition) calculated by the formula: Q = 0.1*N/(0.1+RMSD)
PLOTS : T - Flat text file (output from LGA program, rotated structure)
PLOTS : P - Plot of superimposed structures (3D plot colored as bars)
Citing LGA:
Zemla A., "LGA - a Method for Finding 3D Similarities in Protein Structures",
Nucleic Acids Research, 2003, Vol. 31, No. 13, pp. 3370-3374.
[MEDLINE]

