LGA
Sequence Independent Analysis (LGA)
Frame of reference: Cat.Q2246_545_225.5wLII_11195_16
Total number of 3D structures: 24
LGA calculations using distance cutoff DIST: 4.0 A
Residues superimposed below 2.00 A: GREEN
Residues superimposed below 4.00 A: YELLOW
Residues superimposed below 6.00 A: ORANGE
Residues superimposed below 8.00 A: BROWN
Residues superimposed above 8.00 A or not aligned: RED
Terminal residues not aligned: GREY
Structure Deviation Summary
Calculations based on one final LGA superposition
(Bar representation of 3D plots, TEXT)
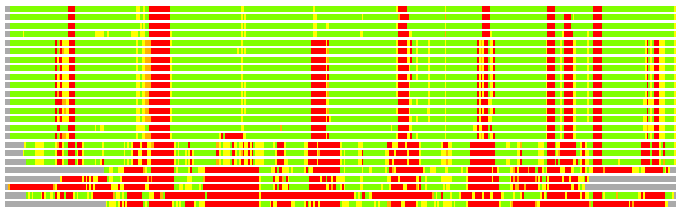
Structures ordered by LGA_S - score
| Structure |
NS |
NT |
N(dist=4.0) |
RMSD(N) |
Seq_ID(N) |
LGA_S |
LGA_Q |
PLOTS |
| 2ixx_A |
354 |
372 |
335 |
0.84 |
11.64 |
88.463 |
35.532 |
T P |
| 2j1n_A |
346 |
372 |
331 |
0.94 |
9.67 |
86.798 |
31.970 |
T P |
| 2ixw_A |
343 |
372 |
331 |
0.93 |
11.18 |
86.729 |
32.009 |
T P |
| 1osm_A |
342 |
372 |
329 |
1.25 |
10.33 |
83.833 |
24.388 |
T P |
| 1gfq_A |
340 |
372 |
316 |
1.35 |
10.76 |
81.350 |
21.738 |
T P |
| 1mpf_A |
340 |
372 |
317 |
1.37 |
11.04 |
81.341 |
21.517 |
T P |
| 1gfm_A |
340 |
372 |
316 |
1.35 |
11.08 |
81.307 |
21.849 |
T P |
| 1gfp_A |
340 |
372 |
316 |
1.35 |
11.08 |
81.296 |
21.803 |
T P |
| 1gfo_A |
340 |
372 |
315 |
1.34 |
11.43 |
81.262 |
21.861 |
T P |
| 2zfg_A |
339 |
372 |
316 |
1.35 |
11.08 |
81.156 |
21.863 |
T P |
| 1hxx_A |
340 |
372 |
314 |
1.31 |
11.15 |
81.041 |
22.256 |
T P |
| 1bt9_A |
340 |
372 |
313 |
1.30 |
11.18 |
81.038 |
22.322 |
T P |
| 1hxt_A |
340 |
372 |
314 |
1.29 |
11.15 |
80.992 |
22.594 |
T P |
| 1hxu_A |
340 |
372 |
315 |
1.39 |
11.43 |
80.853 |
21.135 |
T P |
| 1pho_A |
330 |
372 |
315 |
1.27 |
10.16 |
80.793 |
23.062 |
T P |
| 1gfn_A |
327 |
372 |
305 |
1.37 |
11.48 |
78.444 |
20.808 |
T P |
| 1e54_A |
332 |
372 |
247 |
1.87 |
7.69 |
51.213 |
12.537 |
T P |
| 2fgr_A |
332 |
372 |
245 |
1.88 |
7.35 |
48.721 |
12.387 |
T P |
| 2fgq_X |
330 |
372 |
244 |
1.90 |
7.38 |
48.468 |
12.192 |
T P |
| 2iwv_A |
277 |
372 |
153 |
2.22 |
4.58 |
29.526 |
6.602 |
T P |
| 3bs0_A |
414 |
372 |
142 |
2.14 |
3.52 |
28.596 |
6.331 |
T P |
| 3bry_A |
389 |
372 |
138 |
2.21 |
5.07 |
26.452 |
5.967 |
T P |
| 2f1c_X |
252 |
372 |
135 |
2.13 |
11.85 |
25.624 |
6.043 |
T P |
| 2jqy_A |
280 |
372 |
123 |
2.24 |
7.32 |
23.858 |
5.258 |
T P |
NS : Total number of residues in Structure (rotated structure)
NT : Total number of residues in TARGET (frame of reference)
N : Total number of residues superimposed under 4.0 Angstrom distance cutoff
RMSD : RMS deviation calculated on all N residues superimposed under 4.0 Angstrom distance cutoff
Seq_Id : Sequence Identity. Percent of identical residues from the total of N aligned.
LGA_S : Structure similarity score calculated by internal LGA procedure (see LGA paper for details)
LGA_Q : Score (how tight is the superposition) calculated by the formula: Q = 0.1*N/(0.1+RMSD)
PLOTS : T - Flat text file (output from LGA program, rotated structure)
PLOTS : P - Plot of superimposed structures (3D plot colored as bars)
Citing LGA:
Zemla A., "LGA - a Method for Finding 3D Similarities in Protein Structures",
Nucleic Acids Research, 2003, Vol. 31, No. 13, pp. 3370-3374.
[MEDLINE]

