LGA
Sequence Independent Analysis (LGA)
Frame of reference: Cat.Q2246_545_240.5wLII_11212_56
Total number of 3D structures: 33
LGA calculations using distance cutoff DIST: 4.0 A
Residues superimposed below 2.00 A: GREEN
Residues superimposed below 4.00 A: YELLOW
Residues superimposed below 6.00 A: ORANGE
Residues superimposed below 8.00 A: BROWN
Residues superimposed above 8.00 A or not aligned: RED
Terminal residues not aligned: GREY
Structure Deviation Summary
Calculations based on one final LGA superposition
(Bar representation of 3D plots, TEXT)
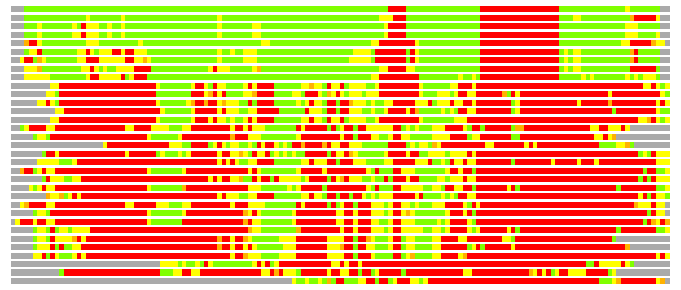
Structures ordered by LGA_S - score
| Structure |
NS |
NT |
N(dist=4.0) |
RMSD(N) |
Seq_ID(N) |
LGA_S |
LGA_Q |
PLOTS |
| 2yr1_A |
257 |
150 |
123 |
0.55 |
19.51 |
81.971 |
18.942 |
T P |
| 1gqn_A |
252 |
150 |
119 |
1.08 |
15.97 |
78.205 |
10.059 |
T P |
| 1sfj_B |
233 |
150 |
122 |
1.24 |
16.39 |
78.135 |
9.092 |
T P |
| 1sfl_B |
233 |
150 |
122 |
1.32 |
16.39 |
77.582 |
8.620 |
T P |
| 2egz_C |
218 |
150 |
113 |
1.58 |
15.93 |
71.515 |
6.708 |
T P |
| 2o7s_A |
500 |
150 |
112 |
1.66 |
18.75 |
69.421 |
6.379 |
T P |
| 2gpt_A |
498 |
150 |
112 |
1.70 |
18.75 |
68.766 |
6.206 |
T P |
| 2ocz_A |
218 |
150 |
111 |
1.74 |
17.12 |
68.468 |
6.039 |
T P |
| 2ox1_A |
196 |
150 |
112 |
1.67 |
16.07 |
68.430 |
6.339 |
T P |
| 1xa3_A |
400 |
150 |
58 |
2.39 |
8.62 |
26.692 |
2.330 |
T P |
| 2g04_A |
353 |
150 |
58 |
2.46 |
8.62 |
26.633 |
2.267 |
T P |
| 1vgq_A |
427 |
150 |
60 |
2.62 |
11.67 |
25.927 |
2.208 |
T P |
| 2vjp_A |
427 |
150 |
57 |
2.30 |
17.54 |
25.648 |
2.379 |
T P |
| 1xk7_A |
407 |
150 |
58 |
2.37 |
6.90 |
25.647 |
2.344 |
T P |
| 2vjm_A |
427 |
150 |
60 |
2.65 |
11.67 |
25.529 |
2.180 |
T P |
| 2vjm_B |
426 |
150 |
52 |
2.32 |
9.62 |
24.891 |
2.146 |
T P |
| 2gci_A |
354 |
150 |
54 |
2.56 |
9.26 |
24.593 |
2.032 |
T P |
| 1xk6_A |
402 |
150 |
58 |
2.50 |
13.79 |
24.575 |
2.232 |
T P |
| 1t3z_A |
427 |
150 |
55 |
2.62 |
9.09 |
24.410 |
2.025 |
T P |
| 1pqy_A |
412 |
150 |
48 |
2.15 |
8.33 |
24.147 |
2.134 |
T P |
| 1xvu_A |
402 |
150 |
54 |
2.48 |
9.26 |
23.912 |
2.096 |
T P |
| 1q7e_A |
410 |
150 |
51 |
2.21 |
7.84 |
23.798 |
2.209 |
T P |
| 1t4c_B |
426 |
150 |
51 |
2.42 |
13.73 |
23.631 |
2.021 |
T P |
| 2vjn_A |
427 |
150 |
55 |
2.63 |
10.91 |
23.557 |
2.017 |
T P |
| 1q6y_A |
417 |
150 |
48 |
2.22 |
6.25 |
23.408 |
2.065 |
T P |
| 1pt7_A |
415 |
150 |
48 |
2.41 |
6.25 |
23.099 |
1.909 |
T P |
| 1vgr_A |
427 |
150 |
51 |
2.43 |
7.84 |
22.336 |
2.013 |
T P |
| 2vjo_A |
427 |
150 |
50 |
2.58 |
14.00 |
22.032 |
1.863 |
T P |
| 1t4c_A |
427 |
150 |
47 |
2.55 |
6.38 |
21.849 |
1.771 |
T P |
| 2vjq_A |
427 |
150 |
45 |
2.53 |
11.11 |
20.490 |
1.713 |
T P |
| 1p5h_A |
427 |
150 |
37 |
2.41 |
10.81 |
17.689 |
1.476 |
T P |
| 1p4t_A |
156 |
150 |
35 |
2.78 |
8.57 |
16.905 |
1.217 |
T P |
| 2f1v_A |
182 |
150 |
29 |
2.39 |
13.79 |
14.363 |
1.166 |
T P |
NS : Total number of residues in Structure (rotated structure)
NT : Total number of residues in TARGET (frame of reference)
N : Total number of residues superimposed under 4.0 Angstrom distance cutoff
RMSD : RMS deviation calculated on all N residues superimposed under 4.0 Angstrom distance cutoff
Seq_Id : Sequence Identity. Percent of identical residues from the total of N aligned.
LGA_S : Structure similarity score calculated by internal LGA procedure (see LGA paper for details)
LGA_Q : Score (how tight is the superposition) calculated by the formula: Q = 0.1*N/(0.1+RMSD)
PLOTS : T - Flat text file (output from LGA program, rotated structure)
PLOTS : P - Plot of superimposed structures (3D plot colored as bars)
Citing LGA:
Zemla A., "LGA - a Method for Finding 3D Similarities in Protein Structures",
Nucleic Acids Research, 2003, Vol. 31, No. 13, pp. 3370-3374.
[MEDLINE]

