LGA
Sequence Independent Analysis (LGA)
Frame of reference: Cat.Q2246_545_245.5wLII_11212_69
Total number of 3D structures: 83
LGA calculations using distance cutoff DIST: 4.0 A
Residues superimposed below 2.00 A: GREEN
Residues superimposed below 4.00 A: YELLOW
Residues superimposed below 6.00 A: ORANGE
Residues superimposed below 8.00 A: BROWN
Residues superimposed above 8.00 A or not aligned: RED
Terminal residues not aligned: GREY
Structure Deviation Summary
Calculations based on one final LGA superposition
(Bar representation of 3D plots, TEXT)
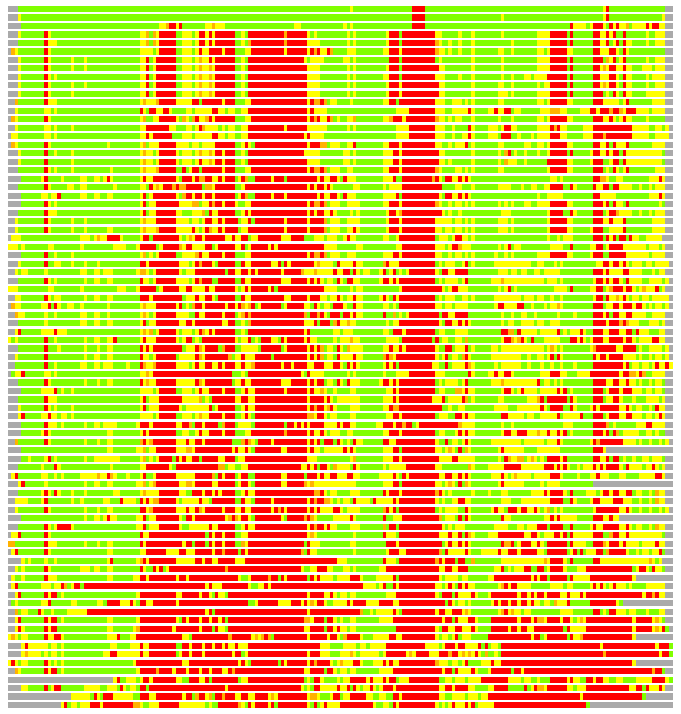
Structures ordered by LGA_S - score
| Structure |
NS |
NT |
N(dist=4.0) |
RMSD(N) |
Seq_ID(N) |
LGA_S |
LGA_Q |
PLOTS |
| 1dts_A |
220 |
202 |
192 |
0.45 |
25.00 |
94.674 |
34.924 |
T P |
| 1byi_A |
224 |
202 |
192 |
0.59 |
25.00 |
94.520 |
27.846 |
T P |
| 2qmo_A |
220 |
202 |
180 |
1.55 |
18.33 |
83.733 |
10.929 |
T P |
| 1jpn_A |
296 |
202 |
141 |
1.85 |
16.31 |
54.800 |
7.234 |
T P |
| 2c03_B |
297 |
202 |
142 |
1.90 |
16.20 |
54.762 |
7.118 |
T P |
| 3e70_C |
307 |
202 |
143 |
2.04 |
10.49 |
53.149 |
6.686 |
T P |
| 2ffh_A |
407 |
202 |
141 |
1.82 |
17.02 |
53.004 |
7.351 |
T P |
| 2j45_A |
297 |
202 |
140 |
1.88 |
16.43 |
52.930 |
7.053 |
T P |
| 2ng1_A |
293 |
202 |
140 |
1.88 |
16.43 |
52.210 |
7.086 |
T P |
| 1ls1_A |
289 |
202 |
140 |
1.89 |
16.43 |
52.146 |
7.042 |
T P |
| 1ffh_A |
287 |
202 |
140 |
1.88 |
16.43 |
51.589 |
7.055 |
T P |
| 1fts_A |
295 |
202 |
140 |
2.09 |
14.29 |
51.260 |
6.397 |
T P |
| 1vcn_A |
506 |
202 |
150 |
2.26 |
16.67 |
50.905 |
6.351 |
T P |
| 3dm9_B |
305 |
202 |
143 |
2.10 |
9.09 |
50.818 |
6.496 |
T P |
| 1s1m_B |
536 |
202 |
149 |
2.23 |
13.42 |
50.711 |
6.408 |
T P |
| 1vco_A |
531 |
202 |
147 |
2.23 |
17.69 |
50.387 |
6.301 |
T P |
| 1vma_A |
294 |
202 |
142 |
2.09 |
11.27 |
50.303 |
6.470 |
T P |
| 1rj9_B |
282 |
202 |
143 |
2.09 |
14.69 |
50.029 |
6.539 |
T P |
| 2j7p_A |
292 |
202 |
142 |
2.07 |
16.20 |
49.839 |
6.553 |
T P |
| 2qy9_A |
300 |
202 |
137 |
1.99 |
13.14 |
49.680 |
6.559 |
T P |
| 2bek_A |
244 |
202 |
136 |
2.11 |
18.38 |
49.627 |
6.155 |
T P |
| 2v3c_C |
403 |
202 |
135 |
2.16 |
12.59 |
49.400 |
5.980 |
T P |
| 1wcv_1 |
243 |
202 |
134 |
2.04 |
16.42 |
49.290 |
6.268 |
T P |
| 2q9c_A |
304 |
202 |
138 |
2.10 |
16.67 |
48.991 |
6.275 |
T P |
| 3dm5_A |
416 |
202 |
137 |
2.01 |
13.14 |
48.834 |
6.490 |
T P |
| 2og2_A |
302 |
202 |
141 |
2.13 |
12.06 |
48.812 |
6.315 |
T P |
| 3b9q_A |
298 |
202 |
141 |
2.17 |
12.06 |
48.720 |
6.223 |
T P |
| 3ea0_B |
242 |
202 |
140 |
2.25 |
10.00 |
48.388 |
5.953 |
T P |
| 1eg7_A |
549 |
202 |
140 |
2.16 |
13.57 |
47.978 |
6.191 |
T P |
| 1okk_D |
265 |
202 |
138 |
2.16 |
16.67 |
47.716 |
6.107 |
T P |
| 1cp2_A |
269 |
202 |
141 |
2.22 |
10.64 |
47.693 |
6.078 |
T P |
| 1ion_A |
243 |
202 |
139 |
2.32 |
15.83 |
47.279 |
5.755 |
T P |
| 1xd8_A |
289 |
202 |
140 |
2.27 |
10.71 |
47.119 |
5.914 |
T P |
| 1fpm_A |
549 |
202 |
136 |
2.17 |
13.24 |
46.891 |
5.988 |
T P |
| 2afh_E |
289 |
202 |
140 |
2.29 |
12.14 |
46.802 |
5.861 |
T P |
| 2ph1_A |
247 |
202 |
138 |
2.24 |
17.39 |
46.718 |
5.887 |
T P |
| 1g3q_A |
237 |
202 |
133 |
2.20 |
15.04 |
46.712 |
5.774 |
T P |
| 1hyq_A |
232 |
202 |
132 |
2.16 |
14.39 |
46.697 |
5.830 |
T P |
| 2j37_W |
479 |
202 |
139 |
2.19 |
10.07 |
46.633 |
6.080 |
T P |
| 1yrb_B |
260 |
202 |
140 |
2.25 |
13.57 |
46.591 |
5.950 |
T P |
| 1g20_H |
262 |
202 |
139 |
2.29 |
12.23 |
46.421 |
5.813 |
T P |
| 2c8v_A |
271 |
202 |
139 |
2.33 |
10.07 |
46.199 |
5.722 |
T P |
| 3end_A |
268 |
202 |
137 |
2.33 |
9.49 |
46.007 |
5.633 |
T P |
| 2vo1_A |
235 |
202 |
130 |
2.14 |
15.38 |
46.006 |
5.816 |
T P |
| 1xdb_A |
289 |
202 |
137 |
2.28 |
10.22 |
45.791 |
5.767 |
T P |
| 2j7p_D |
265 |
202 |
138 |
2.26 |
16.67 |
45.754 |
5.858 |
T P |
| 1zu4_A |
305 |
202 |
132 |
2.13 |
11.36 |
45.692 |
5.906 |
T P |
| 1qzx_A |
425 |
202 |
133 |
2.19 |
9.02 |
45.547 |
5.812 |
T P |
| 1rj9_A |
277 |
202 |
136 |
2.28 |
16.18 |
45.176 |
5.721 |
T P |
| 1ihu_A |
540 |
202 |
129 |
2.19 |
17.83 |
45.043 |
5.642 |
T P |
| 2iyl_D |
271 |
202 |
132 |
2.22 |
16.67 |
44.724 |
5.686 |
T P |
| 1rw4_A |
271 |
202 |
132 |
2.28 |
10.61 |
44.681 |
5.539 |
T P |
| 2ved_B |
254 |
202 |
125 |
2.14 |
8.00 |
44.500 |
5.581 |
T P |
| 2px0_A |
258 |
202 |
134 |
2.24 |
11.94 |
44.388 |
5.724 |
T P |
| 2c5m_C |
252 |
202 |
128 |
2.23 |
14.84 |
44.374 |
5.499 |
T P |
| 2oze_A |
284 |
202 |
136 |
2.43 |
13.24 |
44.163 |
5.369 |
T P |
| 3bfv_A |
241 |
202 |
128 |
2.25 |
9.38 |
44.161 |
5.443 |
T P |
| 2j28_9 |
430 |
202 |
132 |
2.26 |
9.85 |
44.115 |
5.590 |
T P |
| 1j8m_F |
295 |
202 |
136 |
2.44 |
8.82 |
44.113 |
5.357 |
T P |
| 1de0_A |
289 |
202 |
131 |
2.19 |
11.45 |
43.903 |
5.710 |
T P |
| 3cio_A |
255 |
202 |
123 |
2.19 |
15.45 |
43.522 |
5.372 |
T P |
| 1j8y_F |
291 |
202 |
128 |
2.17 |
8.59 |
43.422 |
5.627 |
T P |
| 2p67_A |
302 |
202 |
119 |
2.10 |
14.29 |
41.141 |
5.405 |
T P |
| 2qm7_A |
314 |
202 |
118 |
2.23 |
14.41 |
41.140 |
5.074 |
T P |
| 2qm8_A |
320 |
202 |
120 |
2.16 |
13.33 |
41.107 |
5.299 |
T P |
| 3cwq_A |
206 |
202 |
123 |
2.35 |
14.63 |
40.762 |
5.030 |
T P |
| 1np6_B |
169 |
202 |
105 |
1.98 |
11.43 |
37.013 |
5.042 |
T P |
| 2yvu_A |
179 |
202 |
97 |
2.27 |
13.40 |
32.909 |
4.098 |
T P |
| 1b23_P |
405 |
202 |
98 |
2.42 |
15.31 |
32.624 |
3.887 |
T P |
| 2gc6_A |
406 |
202 |
90 |
2.20 |
12.22 |
32.116 |
3.918 |
T P |
| 2p5t_D |
247 |
202 |
97 |
2.33 |
12.37 |
31.608 |
3.988 |
T P |
| 1d2e_A |
397 |
202 |
96 |
2.39 |
16.67 |
31.585 |
3.863 |
T P |
| 2gc5_A |
407 |
202 |
89 |
2.35 |
12.36 |
31.536 |
3.631 |
T P |
| 1jbw_A |
409 |
202 |
88 |
2.23 |
11.36 |
30.800 |
3.772 |
T P |
| 3cr8_A |
493 |
202 |
84 |
2.46 |
9.52 |
30.077 |
3.276 |
T P |
| 2reb_A |
303 |
202 |
87 |
2.57 |
14.94 |
29.557 |
3.254 |
T P |
| 1u94_A |
306 |
202 |
86 |
2.64 |
13.95 |
29.545 |
3.136 |
T P |
| 2gks_B |
529 |
202 |
82 |
2.28 |
10.98 |
29.234 |
3.450 |
T P |
| 2gcb_A |
407 |
202 |
79 |
2.13 |
13.92 |
29.211 |
3.538 |
T P |
| 2dy1_A |
660 |
202 |
89 |
2.50 |
6.74 |
28.746 |
3.417 |
T P |
| 2f1r_A |
148 |
202 |
88 |
2.46 |
12.50 |
28.520 |
3.433 |
T P |
| 2fn8_A |
292 |
202 |
70 |
2.85 |
4.29 |
22.413 |
2.375 |
T P |
| 2fn9_A |
280 |
202 |
66 |
2.75 |
10.61 |
22.082 |
2.317 |
T P |
NS : Total number of residues in Structure (rotated structure)
NT : Total number of residues in TARGET (frame of reference)
N : Total number of residues superimposed under 4.0 Angstrom distance cutoff
RMSD : RMS deviation calculated on all N residues superimposed under 4.0 Angstrom distance cutoff
Seq_Id : Sequence Identity. Percent of identical residues from the total of N aligned.
LGA_S : Structure similarity score calculated by internal LGA procedure (see LGA paper for details)
LGA_Q : Score (how tight is the superposition) calculated by the formula: Q = 0.1*N/(0.1+RMSD)
PLOTS : T - Flat text file (output from LGA program, rotated structure)
PLOTS : P - Plot of superimposed structures (3D plot colored as bars)
Citing LGA:
Zemla A., "LGA - a Method for Finding 3D Similarities in Protein Structures",
Nucleic Acids Research, 2003, Vol. 31, No. 13, pp. 3370-3374.
[MEDLINE]

