LGA
Sequence Independent Analysis (LGA)
Frame of reference: Cat.Q2246_545_343.5wLII_11276_129
Total number of 3D structures: 34
LGA calculations using distance cutoff DIST: 4.0 A
Residues superimposed below 2.00 A: GREEN
Residues superimposed below 4.00 A: YELLOW
Residues superimposed below 6.00 A: ORANGE
Residues superimposed below 8.00 A: BROWN
Residues superimposed above 8.00 A or not aligned: RED
Terminal residues not aligned: GREY
Structure Deviation Summary
Calculations based on one final LGA superposition
(Bar representation of 3D plots, TEXT)
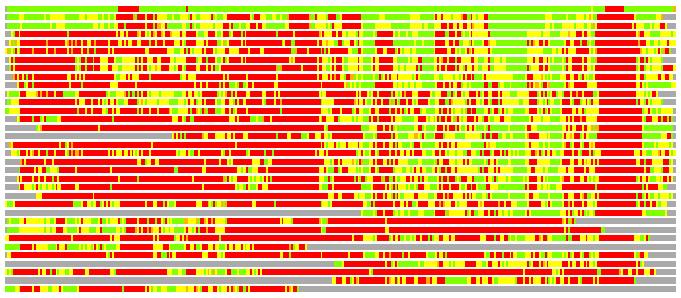
Structures ordered by LGA_S - score
| Structure |
NS |
NT |
N(dist=4.0) |
RMSD(N) |
Seq_ID(N) |
LGA_S |
LGA_Q |
PLOTS |
| 1psw_A |
331 |
344 |
321 |
0.53 |
21.50 |
92.847 |
50.672 |
T P |
| 2gt1_A |
323 |
344 |
252 |
2.11 |
23.81 |
53.931 |
11.405 |
T P |
| 2h1h_A |
322 |
344 |
257 |
2.18 |
23.35 |
52.139 |
11.259 |
T P |
| 3dzc_A |
373 |
344 |
169 |
2.42 |
8.28 |
33.705 |
6.710 |
T P |
| 1o6c_B |
371 |
344 |
164 |
2.44 |
14.02 |
32.840 |
6.469 |
T P |
| 1v4v_A |
373 |
344 |
168 |
2.59 |
14.88 |
31.919 |
6.238 |
T P |
| 1f6d_A |
376 |
344 |
167 |
2.48 |
8.98 |
31.153 |
6.470 |
T P |
| 1vgv_A |
376 |
344 |
165 |
2.49 |
9.09 |
31.047 |
6.378 |
T P |
| 3beo_A |
375 |
344 |
163 |
2.63 |
10.43 |
30.001 |
5.970 |
T P |
| 1iir_A |
382 |
344 |
143 |
2.38 |
9.09 |
29.527 |
5.762 |
T P |
| 3c4v_A |
393 |
344 |
156 |
2.60 |
14.10 |
29.484 |
5.769 |
T P |
| 2gek_A |
361 |
344 |
168 |
2.63 |
8.93 |
28.894 |
6.149 |
T P |
| 2jjm_A |
359 |
344 |
153 |
2.57 |
9.15 |
27.884 |
5.741 |
T P |
| 2vsn_A |
534 |
344 |
143 |
2.51 |
10.49 |
26.685 |
5.476 |
T P |
| 2f9f_A |
166 |
344 |
116 |
2.05 |
9.48 |
26.628 |
5.384 |
T P |
| 1f0k_A |
351 |
344 |
137 |
2.36 |
8.76 |
26.565 |
5.563 |
T P |
| 3c48_A |
399 |
344 |
135 |
2.40 |
11.11 |
26.339 |
5.404 |
T P |
| 2iw1_A |
370 |
344 |
145 |
2.75 |
9.66 |
26.111 |
5.092 |
T P |
| 1rzu_B |
478 |
344 |
143 |
2.59 |
10.49 |
25.839 |
5.313 |
T P |
| 2vsy_A |
547 |
344 |
142 |
2.61 |
7.75 |
25.399 |
5.237 |
T P |
| 2bis_A |
440 |
344 |
133 |
2.46 |
5.26 |
24.877 |
5.192 |
T P |
| 2iuy_A |
340 |
344 |
122 |
2.44 |
9.02 |
23.578 |
4.801 |
T P |
| 2bfw_A |
196 |
344 |
122 |
2.46 |
6.56 |
23.217 |
4.757 |
T P |
| 2iv3_C |
340 |
344 |
123 |
2.60 |
12.20 |
22.673 |
4.563 |
T P |
| 1rcu_A |
170 |
344 |
108 |
2.16 |
18.52 |
21.727 |
4.785 |
T P |
| 3clr_D |
319 |
344 |
100 |
2.59 |
11.00 |
19.972 |
3.723 |
T P |
| 3cls_D |
318 |
344 |
96 |
2.48 |
12.50 |
19.473 |
3.725 |
T P |
| 3clt_D |
319 |
344 |
96 |
2.56 |
5.21 |
17.972 |
3.609 |
T P |
| 1gpj_A |
399 |
344 |
91 |
2.38 |
16.48 |
17.507 |
3.671 |
T P |
| 3clu_D |
319 |
344 |
90 |
2.52 |
3.33 |
17.167 |
3.438 |
T P |
| 1pvo_A |
408 |
344 |
89 |
2.66 |
10.11 |
16.310 |
3.221 |
T P |
| 1jf9_A |
405 |
344 |
81 |
2.56 |
7.41 |
15.085 |
3.050 |
T P |
| 2ht1_A |
324 |
344 |
73 |
2.76 |
10.96 |
14.126 |
2.550 |
T P |
| 3e70_C |
307 |
344 |
75 |
2.50 |
6.67 |
14.107 |
2.888 |
T P |
NS : Total number of residues in Structure (rotated structure)
NT : Total number of residues in TARGET (frame of reference)
N : Total number of residues superimposed under 4.0 Angstrom distance cutoff
RMSD : RMS deviation calculated on all N residues superimposed under 4.0 Angstrom distance cutoff
Seq_Id : Sequence Identity. Percent of identical residues from the total of N aligned.
LGA_S : Structure similarity score calculated by internal LGA procedure (see LGA paper for details)
LGA_Q : Score (how tight is the superposition) calculated by the formula: Q = 0.1*N/(0.1+RMSD)
PLOTS : T - Flat text file (output from LGA program, rotated structure)
PLOTS : P - Plot of superimposed structures (3D plot colored as bars)
Citing LGA:
Zemla A., "LGA - a Method for Finding 3D Similarities in Protein Structures",
Nucleic Acids Research, 2003, Vol. 31, No. 13, pp. 3370-3374.
[MEDLINE]

