LGA
Sequence Independent Analysis (LGA)
Frame of reference: Cat.Q2246_545_349.5wLII_11276_162
Total number of 3D structures: 13
LGA calculations using distance cutoff DIST: 4.0 A
Residues superimposed below 2.00 A: GREEN
Residues superimposed below 4.00 A: YELLOW
Residues superimposed below 6.00 A: ORANGE
Residues superimposed below 8.00 A: BROWN
Residues superimposed above 8.00 A or not aligned: RED
Terminal residues not aligned: GREY
Structure Deviation Summary
Calculations based on one final LGA superposition
(Bar representation of 3D plots, TEXT)
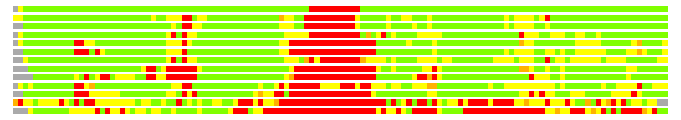
Structures ordered by LGA_S - score
| Structure |
NS |
NT |
N(dist=4.0) |
RMSD(N) |
Seq_ID(N) |
LGA_S |
LGA_Q |
PLOTS |
| 1jx7_A |
117 |
128 |
117 |
0.34 |
22.22 |
91.283 |
26.713 |
T P |
| 2d1p_A |
130 |
128 |
115 |
1.51 |
20.00 |
83.690 |
7.141 |
T P |
| 2hy5_A |
130 |
128 |
114 |
1.39 |
14.91 |
83.402 |
7.639 |
T P |
| 2hy5_B |
132 |
128 |
113 |
1.76 |
20.35 |
79.910 |
6.059 |
T P |
| 1l1s_A |
111 |
128 |
107 |
1.74 |
22.43 |
76.491 |
5.816 |
T P |
| 2pd2_A |
108 |
128 |
104 |
1.67 |
18.27 |
74.603 |
5.875 |
T P |
| 2d1p_B |
119 |
128 |
114 |
1.98 |
13.16 |
73.715 |
5.485 |
T P |
| 2hy5_C |
101 |
128 |
100 |
1.60 |
22.00 |
71.344 |
5.871 |
T P |
| 2d1p_C |
95 |
128 |
93 |
1.59 |
20.43 |
67.373 |
5.517 |
T P |
| 2qs7_A |
138 |
128 |
110 |
2.09 |
11.82 |
62.837 |
5.013 |
T P |
| 2fb6_A |
116 |
128 |
100 |
1.86 |
11.00 |
61.980 |
5.099 |
T P |
| 2a3l_A |
616 |
128 |
76 |
2.64 |
7.89 |
37.660 |
2.776 |
T P |
| 2nvv_A |
496 |
128 |
64 |
2.43 |
10.94 |
34.951 |
2.534 |
T P |
NS : Total number of residues in Structure (rotated structure)
NT : Total number of residues in TARGET (frame of reference)
N : Total number of residues superimposed under 4.0 Angstrom distance cutoff
RMSD : RMS deviation calculated on all N residues superimposed under 4.0 Angstrom distance cutoff
Seq_Id : Sequence Identity. Percent of identical residues from the total of N aligned.
LGA_S : Structure similarity score calculated by internal LGA procedure (see LGA paper for details)
LGA_Q : Score (how tight is the superposition) calculated by the formula: Q = 0.1*N/(0.1+RMSD)
PLOTS : T - Flat text file (output from LGA program, rotated structure)
PLOTS : P - Plot of superimposed structures (3D plot colored as bars)
Citing LGA:
Zemla A., "LGA - a Method for Finding 3D Similarities in Protein Structures",
Nucleic Acids Research, 2003, Vol. 31, No. 13, pp. 3370-3374.
[MEDLINE]

