LGA
Sequence Independent Analysis (LGA)
Frame of reference: Cat.Q2246_545_373.5wLII_11277_57
Total number of 3D structures: 46
LGA calculations using distance cutoff DIST: 4.0 A
Residues superimposed below 2.00 A: GREEN
Residues superimposed below 4.00 A: YELLOW
Residues superimposed below 6.00 A: ORANGE
Residues superimposed below 8.00 A: BROWN
Residues superimposed above 8.00 A or not aligned: RED
Terminal residues not aligned: GREY
Structure Deviation Summary
Calculations based on one final LGA superposition
(Bar representation of 3D plots, TEXT)
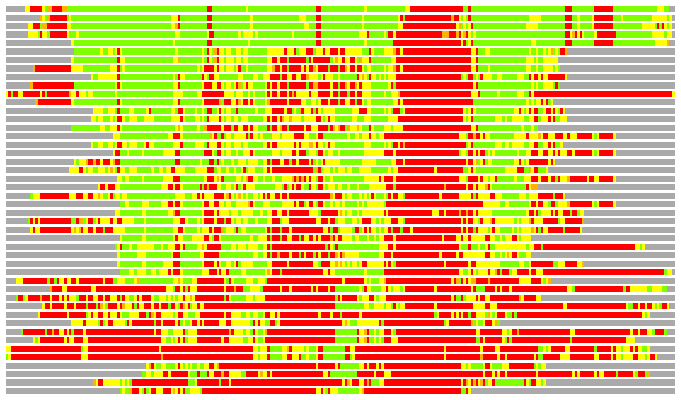
Structures ordered by LGA_S - score
| Structure |
NS |
NT |
N(dist=4.0) |
RMSD(N) |
Seq_ID(N) |
LGA_S |
LGA_Q |
PLOTS |
| 1gpm_A |
501 |
276 |
219 |
1.02 |
19.18 |
78.413 |
19.543 |
T P |
| 2ywc_A |
475 |
276 |
215 |
1.44 |
18.60 |
74.021 |
13.932 |
T P |
| 2ywb_A |
471 |
276 |
214 |
1.45 |
17.76 |
72.971 |
13.767 |
T P |
| 2vxo_B |
658 |
276 |
215 |
1.60 |
15.35 |
72.846 |
12.611 |
T P |
| 2dpl_A |
287 |
276 |
202 |
1.17 |
16.83 |
70.417 |
15.951 |
T P |
| 1xng_B |
262 |
276 |
156 |
2.15 |
12.18 |
41.984 |
6.945 |
T P |
| 3fiu_A |
238 |
276 |
142 |
2.03 |
12.68 |
39.351 |
6.674 |
T P |
| 2e18_A |
256 |
276 |
140 |
2.06 |
16.43 |
38.663 |
6.486 |
T P |
| 1ni5_A |
433 |
276 |
138 |
2.20 |
21.01 |
37.320 |
5.992 |
T P |
| 1kqp_A |
271 |
276 |
131 |
1.92 |
15.27 |
36.663 |
6.472 |
T P |
| 3dla_C |
657 |
276 |
135 |
2.17 |
12.59 |
36.279 |
5.948 |
T P |
| 3dpi_A |
269 |
276 |
129 |
1.91 |
17.83 |
36.260 |
6.429 |
T P |
| 2goy_B |
223 |
276 |
138 |
2.17 |
21.01 |
35.075 |
6.069 |
T P |
| 2o8v_A |
229 |
276 |
153 |
2.34 |
15.69 |
35.029 |
6.275 |
T P |
| 2pzb_A |
250 |
276 |
128 |
2.14 |
12.50 |
34.985 |
5.712 |
T P |
| 1k92_A |
444 |
276 |
139 |
2.31 |
15.83 |
34.877 |
5.760 |
T P |
| 1sur_A |
215 |
276 |
148 |
2.33 |
16.89 |
34.372 |
6.090 |
T P |
| 2nz2_A |
402 |
276 |
134 |
2.28 |
16.42 |
33.152 |
5.624 |
T P |
| 1wy5_A |
311 |
276 |
130 |
2.15 |
21.54 |
33.095 |
5.771 |
T P |
| 1zun_A |
204 |
276 |
138 |
2.31 |
18.12 |
33.020 |
5.717 |
T P |
| 1vl2_A |
398 |
276 |
130 |
2.29 |
19.23 |
32.820 |
5.439 |
T P |
| 2c5s_A |
372 |
276 |
123 |
2.28 |
16.26 |
31.896 |
5.157 |
T P |
| 1q15_A |
491 |
276 |
132 |
2.44 |
15.91 |
30.682 |
5.194 |
T P |
| 1kor_B |
386 |
276 |
128 |
2.39 |
26.56 |
30.588 |
5.143 |
T P |
| 2hma_A |
364 |
276 |
127 |
2.38 |
21.26 |
30.053 |
5.124 |
T P |
| 1ct9_A |
497 |
276 |
129 |
2.57 |
14.73 |
29.872 |
4.823 |
T P |
| 1jgt_B |
500 |
276 |
125 |
2.39 |
14.40 |
29.809 |
5.013 |
T P |
| 2pg3_A |
221 |
276 |
111 |
2.36 |
21.62 |
28.902 |
4.506 |
T P |
| 2d13_A |
215 |
276 |
113 |
2.25 |
17.70 |
28.782 |
4.815 |
T P |
| 3bl5_E |
199 |
276 |
107 |
2.24 |
16.82 |
28.299 |
4.582 |
T P |
| 2der_B |
359 |
276 |
115 |
2.40 |
21.74 |
28.180 |
4.592 |
T P |
| 1ru8_A |
227 |
276 |
104 |
2.22 |
18.27 |
26.210 |
4.479 |
T P |
| 1xa0_A |
318 |
276 |
90 |
2.53 |
10.00 |
21.736 |
3.419 |
T P |
| 1kkj_A |
405 |
276 |
84 |
2.59 |
3.57 |
20.002 |
3.124 |
T P |
| 1o89_A |
320 |
276 |
85 |
2.61 |
7.06 |
19.125 |
3.132 |
T P |
| 1dfo_B |
417 |
276 |
78 |
2.59 |
10.26 |
19.113 |
2.900 |
T P |
| 1knw_A |
421 |
276 |
80 |
2.68 |
8.75 |
18.782 |
2.880 |
T P |
| 3e15_B |
295 |
276 |
76 |
2.70 |
6.58 |
17.393 |
2.715 |
T P |
| 3cog_D |
392 |
276 |
72 |
2.61 |
6.94 |
17.053 |
2.658 |
T P |
| 3elp_B |
343 |
276 |
68 |
2.48 |
13.24 |
16.771 |
2.635 |
T P |
| 1l8a_A |
801 |
276 |
67 |
2.51 |
7.46 |
16.404 |
2.566 |
T P |
| 2qtc_A |
801 |
276 |
67 |
2.51 |
8.96 |
16.316 |
2.571 |
T P |
| 1xrj_B |
213 |
276 |
64 |
2.44 |
4.69 |
16.054 |
2.524 |
T P |
| 1wqa_A |
455 |
276 |
64 |
2.47 |
7.81 |
15.931 |
2.488 |
T P |
| 1uj2_A |
213 |
276 |
58 |
2.67 |
6.90 |
14.348 |
2.094 |
T P |
| 2vpi_A |
187 |
276 |
44 |
2.44 |
6.82 |
11.148 |
1.731 |
T P |
NS : Total number of residues in Structure (rotated structure)
NT : Total number of residues in TARGET (frame of reference)
N : Total number of residues superimposed under 4.0 Angstrom distance cutoff
RMSD : RMS deviation calculated on all N residues superimposed under 4.0 Angstrom distance cutoff
Seq_Id : Sequence Identity. Percent of identical residues from the total of N aligned.
LGA_S : Structure similarity score calculated by internal LGA procedure (see LGA paper for details)
LGA_Q : Score (how tight is the superposition) calculated by the formula: Q = 0.1*N/(0.1+RMSD)
PLOTS : T - Flat text file (output from LGA program, rotated structure)
PLOTS : P - Plot of superimposed structures (3D plot colored as bars)
Citing LGA:
Zemla A., "LGA - a Method for Finding 3D Similarities in Protein Structures",
Nucleic Acids Research, 2003, Vol. 31, No. 13, pp. 3370-3374.
[MEDLINE]

