LGA
Sequence Independent Analysis (LGA)
Frame of reference: Cat.Q2246_545_381.5wLII_11277_84
Total number of 3D structures: 46
LGA calculations using distance cutoff DIST: 4.0 A
Residues superimposed below 2.00 A: GREEN
Residues superimposed below 4.00 A: YELLOW
Residues superimposed below 6.00 A: ORANGE
Residues superimposed below 8.00 A: BROWN
Residues superimposed above 8.00 A or not aligned: RED
Terminal residues not aligned: GREY
Structure Deviation Summary
Calculations based on one final LGA superposition
(Bar representation of 3D plots, TEXT)
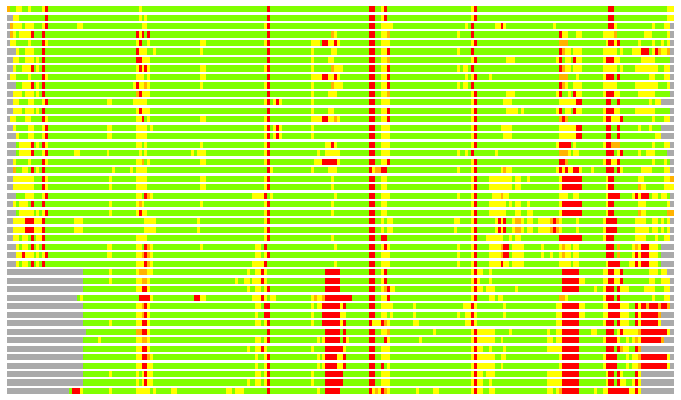
Structures ordered by LGA_S - score
| Structure |
NS |
NT |
N(dist=4.0) |
RMSD(N) |
Seq_ID(N) |
LGA_S |
LGA_Q |
PLOTS |
| 1j2q_A |
237 |
228 |
220 |
0.88 |
21.82 |
94.947 |
22.429 |
T P |
| 1j2p_A |
243 |
228 |
218 |
0.65 |
21.10 |
94.529 |
28.915 |
T P |
| 1iru_A |
244 |
228 |
218 |
1.29 |
15.60 |
90.682 |
15.632 |
T P |
| 1iru_C |
250 |
228 |
216 |
1.42 |
18.98 |
89.874 |
14.251 |
T P |
| 1iru_E |
234 |
228 |
214 |
1.47 |
21.50 |
88.783 |
13.602 |
T P |
| 3bdm_B |
244 |
228 |
212 |
1.50 |
21.23 |
88.348 |
13.281 |
T P |
| 1ryp_D |
241 |
228 |
213 |
1.49 |
18.78 |
88.148 |
13.361 |
T P |
| 1ryp_C |
244 |
228 |
215 |
1.61 |
20.93 |
88.056 |
12.592 |
T P |
| 3bdm_D |
242 |
228 |
213 |
1.52 |
20.66 |
87.952 |
13.113 |
T P |
| 1g0u_B |
235 |
228 |
215 |
1.56 |
20.93 |
87.948 |
12.913 |
T P |
| 1g0u_C |
238 |
228 |
214 |
1.53 |
18.69 |
87.933 |
13.138 |
T P |
| 1ya7_A |
227 |
228 |
211 |
1.32 |
19.91 |
87.916 |
14.831 |
T P |
| 3bdm_C |
241 |
228 |
213 |
1.52 |
18.78 |
87.894 |
13.157 |
T P |
| 1ryp_E |
242 |
228 |
213 |
1.53 |
20.19 |
87.804 |
13.074 |
T P |
| 1yau_A |
222 |
228 |
211 |
1.35 |
19.91 |
87.790 |
14.560 |
T P |
| 1yar_A |
221 |
228 |
210 |
1.30 |
20.00 |
87.696 |
15.042 |
T P |
| 1ryp_B |
250 |
228 |
211 |
1.47 |
19.43 |
87.592 |
13.453 |
T P |
| 1iru_B |
233 |
228 |
213 |
1.50 |
19.72 |
87.298 |
13.327 |
T P |
| 1g0u_D |
230 |
228 |
209 |
1.38 |
19.62 |
87.015 |
14.123 |
T P |
| 1iru_D |
243 |
228 |
210 |
1.55 |
17.14 |
86.721 |
12.717 |
T P |
| 1g0u_F |
242 |
228 |
212 |
1.50 |
15.57 |
86.629 |
13.230 |
T P |
| 1ryp_G |
244 |
228 |
212 |
1.52 |
15.57 |
86.307 |
13.072 |
T P |
| 1iru_F |
238 |
228 |
210 |
1.51 |
17.62 |
86.257 |
13.010 |
T P |
| 3bdm_F |
244 |
228 |
210 |
1.47 |
14.76 |
86.067 |
13.400 |
T P |
| 1z7q_G |
243 |
228 |
210 |
1.57 |
15.71 |
85.796 |
12.612 |
T P |
| 1g0u_G |
240 |
228 |
210 |
1.65 |
13.81 |
85.436 |
12.028 |
T P |
| 1ryp_A |
243 |
228 |
210 |
1.66 |
13.81 |
85.374 |
11.906 |
T P |
| 1iru_G |
245 |
228 |
206 |
1.50 |
11.17 |
85.217 |
12.842 |
T P |
| 1ryp_F |
233 |
228 |
205 |
1.67 |
14.63 |
83.447 |
11.586 |
T P |
| 3bdm_E |
233 |
228 |
205 |
1.67 |
16.10 |
83.174 |
11.590 |
T P |
| 1g0u_E |
230 |
228 |
205 |
1.62 |
14.63 |
83.145 |
11.931 |
T P |
| 1pma_B |
203 |
228 |
181 |
1.52 |
14.92 |
74.763 |
11.191 |
T P |
| 1yar_H |
203 |
228 |
182 |
1.58 |
14.29 |
74.513 |
10.828 |
T P |
| 1j2q_H |
202 |
228 |
179 |
1.57 |
24.02 |
73.634 |
10.705 |
T P |
| 2z5c_C |
189 |
228 |
179 |
1.57 |
18.44 |
73.115 |
10.701 |
T P |
| 1ryp_H |
205 |
228 |
170 |
1.66 |
17.65 |
70.054 |
9.633 |
T P |
| 1g65_N |
196 |
228 |
169 |
1.58 |
17.75 |
69.862 |
10.069 |
T P |
| 1iru_H |
202 |
228 |
168 |
1.58 |
17.86 |
69.720 |
10.002 |
T P |
| 1g65_K |
211 |
228 |
169 |
1.67 |
13.02 |
69.687 |
9.526 |
T P |
| 1iru_L |
201 |
228 |
171 |
1.73 |
12.87 |
69.635 |
9.362 |
T P |
| 1iru_I |
220 |
228 |
168 |
1.55 |
17.26 |
69.493 |
10.207 |
T P |
| 1ryp_L |
212 |
228 |
171 |
1.70 |
12.28 |
69.395 |
9.486 |
T P |
| 1g0u_K |
212 |
228 |
170 |
1.66 |
13.53 |
69.248 |
9.680 |
T P |
| 1ryp_I |
222 |
228 |
170 |
1.53 |
15.88 |
69.131 |
10.401 |
T P |
| 3bdm_H |
222 |
228 |
170 |
1.55 |
15.88 |
68.952 |
10.308 |
T P |
| 1iru_J |
204 |
228 |
170 |
1.75 |
12.35 |
68.652 |
9.202 |
T P |
NS : Total number of residues in Structure (rotated structure)
NT : Total number of residues in TARGET (frame of reference)
N : Total number of residues superimposed under 4.0 Angstrom distance cutoff
RMSD : RMS deviation calculated on all N residues superimposed under 4.0 Angstrom distance cutoff
Seq_Id : Sequence Identity. Percent of identical residues from the total of N aligned.
LGA_S : Structure similarity score calculated by internal LGA procedure (see LGA paper for details)
LGA_Q : Score (how tight is the superposition) calculated by the formula: Q = 0.1*N/(0.1+RMSD)
PLOTS : T - Flat text file (output from LGA program, rotated structure)
PLOTS : P - Plot of superimposed structures (3D plot colored as bars)
Citing LGA:
Zemla A., "LGA - a Method for Finding 3D Similarities in Protein Structures",
Nucleic Acids Research, 2003, Vol. 31, No. 13, pp. 3370-3374.
[MEDLINE]

