LGA
Sequence Independent Analysis (LGA)
Frame of reference: Cat.Q2246_545_400.5wLII_11277_136
Total number of 3D structures: 23
LGA calculations using distance cutoff DIST: 4.0 A
Residues superimposed below 2.00 A: GREEN
Residues superimposed below 4.00 A: YELLOW
Residues superimposed below 6.00 A: ORANGE
Residues superimposed below 8.00 A: BROWN
Residues superimposed above 8.00 A or not aligned: RED
Terminal residues not aligned: GREY
Structure Deviation Summary
Calculations based on one final LGA superposition
(Bar representation of 3D plots, TEXT)
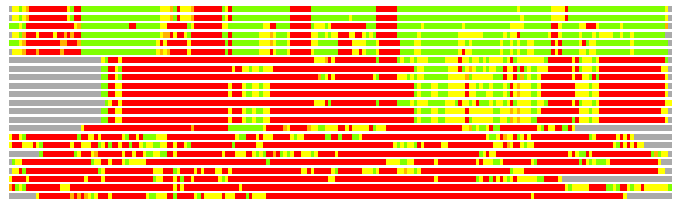
Structures ordered by LGA_S - score
| Structure |
NS |
NT |
N(dist=4.0) |
RMSD(N) |
Seq_ID(N) |
LGA_S |
LGA_Q |
PLOTS |
| 2vrq_A |
493 |
193 |
155 |
1.38 |
14.84 |
78.654 |
10.445 |
T P |
| 2vrk_A |
493 |
193 |
155 |
1.41 |
14.84 |
78.472 |
10.237 |
T P |
| 3fw6_A |
519 |
193 |
133 |
1.65 |
11.28 |
63.901 |
7.611 |
T P |
| 2c7f_D |
499 |
193 |
145 |
2.03 |
6.90 |
60.769 |
6.802 |
T P |
| 1qw9_A |
497 |
193 |
142 |
1.83 |
9.15 |
60.701 |
7.359 |
T P |
| 1pz3_A |
497 |
193 |
140 |
1.90 |
8.57 |
60.223 |
7.006 |
T P |
| 1zj5_A |
232 |
193 |
59 |
2.47 |
8.47 |
20.113 |
2.296 |
T P |
| 1zi8_A |
233 |
193 |
58 |
2.46 |
8.62 |
19.706 |
2.266 |
T P |
| 1zi9_A |
233 |
193 |
56 |
2.41 |
10.71 |
19.692 |
2.228 |
T P |
| 1zic_A |
233 |
193 |
56 |
2.40 |
8.93 |
19.534 |
2.244 |
T P |
| 1zix_A |
233 |
193 |
56 |
2.43 |
8.93 |
19.245 |
2.218 |
T P |
| 1zi6_A |
233 |
193 |
58 |
2.45 |
10.34 |
19.207 |
2.278 |
T P |
| 1zj4_A |
232 |
193 |
57 |
2.54 |
8.77 |
19.064 |
2.156 |
T P |
| 1din_A |
233 |
193 |
56 |
2.50 |
8.93 |
19.064 |
2.157 |
T P |
| 1rh1_A |
490 |
193 |
51 |
2.50 |
1.96 |
18.564 |
1.963 |
T P |
| 2cjp_A |
320 |
193 |
49 |
2.58 |
2.04 |
17.857 |
1.830 |
T P |
| 1ggv_A |
231 |
193 |
54 |
2.79 |
3.70 |
16.964 |
1.866 |
T P |
| 3cxu_A |
320 |
193 |
48 |
2.66 |
12.50 |
16.572 |
1.737 |
T P |
| 3f67_A |
240 |
193 |
42 |
2.76 |
4.76 |
14.082 |
1.467 |
T P |
| 2b9v_A |
608 |
193 |
38 |
2.86 |
5.26 |
12.458 |
1.285 |
T P |
| 1ryy_A |
612 |
193 |
32 |
2.70 |
0.00 |
11.283 |
1.141 |
T P |
| 1nx9_A |
617 |
193 |
33 |
2.65 |
3.03 |
11.191 |
1.198 |
T P |
| 1c4x_A |
281 |
193 |
32 |
3.05 |
15.62 |
10.655 |
1.015 |
T P |
NS : Total number of residues in Structure (rotated structure)
NT : Total number of residues in TARGET (frame of reference)
N : Total number of residues superimposed under 4.0 Angstrom distance cutoff
RMSD : RMS deviation calculated on all N residues superimposed under 4.0 Angstrom distance cutoff
Seq_Id : Sequence Identity. Percent of identical residues from the total of N aligned.
LGA_S : Structure similarity score calculated by internal LGA procedure (see LGA paper for details)
LGA_Q : Score (how tight is the superposition) calculated by the formula: Q = 0.1*N/(0.1+RMSD)
PLOTS : T - Flat text file (output from LGA program, rotated structure)
PLOTS : P - Plot of superimposed structures (3D plot colored as bars)
Citing LGA:
Zemla A., "LGA - a Method for Finding 3D Similarities in Protein Structures",
Nucleic Acids Research, 2003, Vol. 31, No. 13, pp. 3370-3374.
[MEDLINE]

