LGA
Sequence Independent Analysis (LGA)
Frame of reference: Cat.Q2246_545_486.5wLII_11346_138
Total number of 3D structures: 70
LGA calculations using distance cutoff DIST: 4.0 A
Residues superimposed below 2.00 A: GREEN
Residues superimposed below 4.00 A: YELLOW
Residues superimposed below 6.00 A: ORANGE
Residues superimposed below 8.00 A: BROWN
Residues superimposed above 8.00 A or not aligned: RED
Terminal residues not aligned: GREY
Structure Deviation Summary
Calculations based on one final LGA superposition
(Bar representation of 3D plots, TEXT)
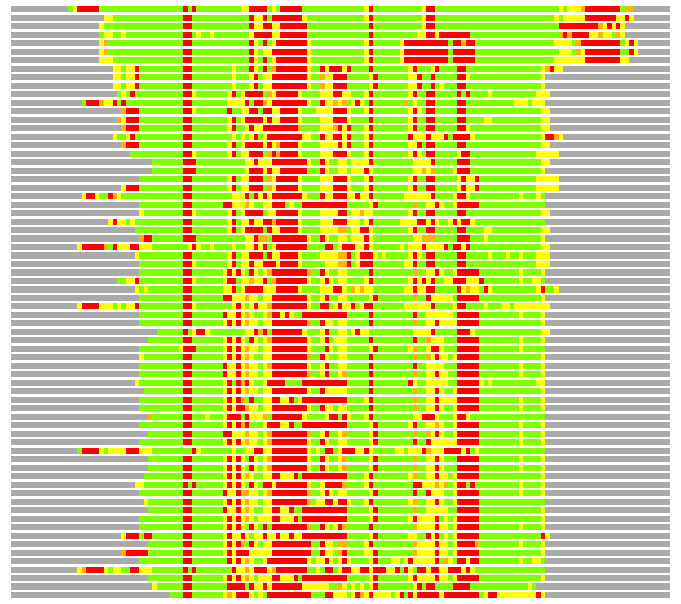
Structures ordered by LGA_S - score
| Structure |
NS |
NT |
N(dist=4.0) |
RMSD(N) |
Seq_ID(N) |
LGA_S |
LGA_Q |
PLOTS |
| 1nir_B |
539 |
149 |
101 |
1.49 |
20.79 |
65.648 |
6.347 |
T P |
| 1e2r_B |
543 |
149 |
98 |
1.38 |
16.33 |
62.416 |
6.608 |
T P |
| 1hzu_A |
521 |
149 |
94 |
1.27 |
19.15 |
62.134 |
6.879 |
T P |
| 1hzv_A |
514 |
149 |
92 |
1.76 |
19.57 |
57.150 |
4.949 |
T P |
| 1gq1_A |
559 |
149 |
86 |
1.61 |
13.95 |
54.963 |
5.015 |
T P |
| 1qks_A |
559 |
149 |
86 |
1.61 |
15.12 |
54.722 |
5.030 |
T P |
| 1hj5_A |
559 |
149 |
85 |
1.57 |
12.94 |
54.159 |
5.081 |
T P |
| 2d0w_A |
168 |
149 |
79 |
1.70 |
16.46 |
48.875 |
4.393 |
T P |
| 2gc4_D |
147 |
149 |
78 |
1.68 |
19.23 |
48.511 |
4.371 |
T P |
| 1mg2_D |
147 |
149 |
78 |
1.70 |
19.23 |
48.148 |
4.334 |
T P |
| 1ls9_A |
91 |
149 |
77 |
1.81 |
19.48 |
46.907 |
4.023 |
T P |
| 1yiq_A |
684 |
149 |
80 |
2.01 |
18.75 |
46.887 |
3.787 |
T P |
| 2zbo_A |
86 |
149 |
74 |
1.77 |
22.97 |
46.220 |
3.956 |
T P |
| 1ctj_A |
89 |
149 |
74 |
1.76 |
21.62 |
46.064 |
3.981 |
T P |
| 1c6r_A |
88 |
149 |
74 |
1.79 |
20.27 |
45.741 |
3.917 |
T P |
| 1umm_A |
149 |
149 |
76 |
1.87 |
18.42 |
45.550 |
3.850 |
T P |
| 1cyj_A |
90 |
149 |
74 |
1.85 |
20.27 |
45.517 |
3.788 |
T P |
| 2dge_A |
102 |
149 |
77 |
1.76 |
18.18 |
43.838 |
4.139 |
T P |
| 1dt1_A |
129 |
149 |
72 |
1.83 |
16.67 |
43.745 |
3.733 |
T P |
| 2fwl_A |
129 |
149 |
71 |
1.83 |
16.90 |
43.548 |
3.672 |
T P |
| 2v07_A |
98 |
149 |
75 |
1.85 |
17.33 |
42.605 |
3.843 |
T P |
| 2ce0_A |
99 |
149 |
76 |
1.87 |
18.42 |
42.548 |
3.863 |
T P |
| 1kv9_A |
664 |
149 |
81 |
1.95 |
16.05 |
41.907 |
3.953 |
T P |
| 2b12_B |
108 |
149 |
68 |
1.85 |
17.65 |
41.780 |
3.479 |
T P |
| 2v08_A |
84 |
149 |
74 |
1.71 |
24.32 |
41.711 |
4.081 |
T P |
| 1f1c_A |
129 |
149 |
77 |
1.97 |
20.78 |
41.549 |
3.718 |
T P |
| 1gdv_A |
85 |
149 |
75 |
1.87 |
25.33 |
41.519 |
3.815 |
T P |
| 1c52_A |
131 |
149 |
73 |
2.11 |
16.44 |
40.368 |
3.310 |
T P |
| 1m70_A |
190 |
149 |
78 |
2.02 |
17.95 |
40.071 |
3.683 |
T P |
| 1kib_A |
89 |
149 |
74 |
1.96 |
24.32 |
39.942 |
3.598 |
T P |
| 1f1f_A |
88 |
149 |
73 |
1.90 |
23.29 |
39.633 |
3.650 |
T P |
| 1csu_A |
108 |
149 |
71 |
1.92 |
21.13 |
39.243 |
3.512 |
T P |
| 3cp5_A |
116 |
149 |
75 |
2.00 |
18.67 |
39.142 |
3.574 |
T P |
| 1c6s_A |
87 |
149 |
76 |
2.10 |
22.37 |
38.873 |
3.453 |
T P |
| 1irv_A |
108 |
149 |
72 |
2.02 |
19.44 |
38.717 |
3.393 |
T P |
| 1kb0_A |
669 |
149 |
81 |
2.29 |
17.28 |
38.532 |
3.394 |
T P |
| 1csv_A |
108 |
149 |
68 |
1.85 |
20.59 |
38.318 |
3.487 |
T P |
| 1i54_A |
103 |
149 |
72 |
1.94 |
22.22 |
38.204 |
3.521 |
T P |
| 1foc_A |
128 |
149 |
71 |
1.89 |
15.49 |
38.179 |
3.559 |
T P |
| 1rap_A |
108 |
149 |
69 |
1.91 |
21.74 |
38.166 |
3.429 |
T P |
| 1co6_A |
107 |
149 |
70 |
2.01 |
15.71 |
38.074 |
3.317 |
T P |
| 2aiu_A |
104 |
149 |
72 |
1.97 |
16.67 |
38.039 |
3.485 |
T P |
| 1raq_A |
108 |
149 |
72 |
1.96 |
20.83 |
38.020 |
3.491 |
T P |
| 1chj_A |
108 |
149 |
72 |
2.00 |
20.83 |
37.901 |
3.423 |
T P |
| 1cyc_A |
103 |
149 |
67 |
1.78 |
20.90 |
37.799 |
3.570 |
T P |
| 1wej_F |
104 |
149 |
71 |
1.96 |
18.31 |
37.717 |
3.447 |
T P |
| 1yeb_A |
108 |
149 |
67 |
1.87 |
19.40 |
37.707 |
3.402 |
T P |
| 1ccr_A |
111 |
149 |
71 |
1.99 |
16.90 |
37.600 |
3.405 |
T P |
| 1wve_C |
75 |
149 |
68 |
2.00 |
22.06 |
37.396 |
3.231 |
T P |
| 2jti_B |
103 |
149 |
66 |
1.83 |
18.18 |
37.275 |
3.421 |
T P |
| 1csw_A |
108 |
149 |
70 |
2.04 |
20.00 |
36.990 |
3.275 |
T P |
| 3cx5_W |
108 |
149 |
71 |
1.96 |
19.72 |
36.861 |
3.453 |
T P |
| 1h1o_A |
172 |
149 |
76 |
2.10 |
22.37 |
36.695 |
3.453 |
T P |
| 1ycc_A |
108 |
149 |
69 |
1.97 |
20.29 |
36.660 |
3.340 |
T P |
| 2jqr_A |
108 |
149 |
69 |
1.97 |
20.29 |
36.660 |
3.340 |
T P |
| 2b4z_A |
104 |
149 |
67 |
1.91 |
16.42 |
36.587 |
3.331 |
T P |
| 1w2l_A |
97 |
149 |
69 |
1.93 |
7.25 |
36.465 |
3.400 |
T P |
| 2ycc_A |
108 |
149 |
71 |
2.01 |
19.72 |
36.341 |
3.366 |
T P |
| 1hro_A |
105 |
149 |
68 |
1.94 |
16.18 |
35.826 |
3.326 |
T P |
| 1ctz_A |
108 |
149 |
67 |
2.02 |
17.91 |
35.696 |
3.167 |
T P |
| 2b11_B |
108 |
149 |
67 |
2.00 |
16.42 |
35.234 |
3.184 |
T P |
| 1chi_A |
108 |
149 |
69 |
1.97 |
20.29 |
34.716 |
3.333 |
T P |
| 1yfc_A |
108 |
149 |
66 |
1.98 |
16.67 |
34.366 |
3.166 |
T P |
| 1irw_A |
108 |
149 |
69 |
2.09 |
20.29 |
33.963 |
3.147 |
T P |
| 1chh_A |
108 |
149 |
67 |
2.05 |
19.40 |
33.397 |
3.116 |
T P |
| 1j3s_A |
104 |
149 |
70 |
2.15 |
17.14 |
33.299 |
3.109 |
T P |
| 1fcd_C |
174 |
149 |
68 |
2.29 |
16.18 |
33.258 |
2.843 |
T P |
| 2bcn_B |
108 |
149 |
65 |
2.01 |
16.92 |
32.335 |
3.077 |
T P |
| 1kx2_A |
81 |
149 |
64 |
1.94 |
20.31 |
31.737 |
3.142 |
T P |
| 1lms_A |
95 |
149 |
48 |
2.37 |
18.75 |
22.814 |
1.941 |
T P |
NS : Total number of residues in Structure (rotated structure)
NT : Total number of residues in TARGET (frame of reference)
N : Total number of residues superimposed under 4.0 Angstrom distance cutoff
RMSD : RMS deviation calculated on all N residues superimposed under 4.0 Angstrom distance cutoff
Seq_Id : Sequence Identity. Percent of identical residues from the total of N aligned.
LGA_S : Structure similarity score calculated by internal LGA procedure (see LGA paper for details)
LGA_Q : Score (how tight is the superposition) calculated by the formula: Q = 0.1*N/(0.1+RMSD)
PLOTS : T - Flat text file (output from LGA program, rotated structure)
PLOTS : P - Plot of superimposed structures (3D plot colored as bars)
Citing LGA:
Zemla A., "LGA - a Method for Finding 3D Similarities in Protein Structures",
Nucleic Acids Research, 2003, Vol. 31, No. 13, pp. 3370-3374.
[MEDLINE]

