LGA
Sequence Independent Analysis (LGA)
Frame of reference: Cat.Q2246_545_527.5wLII_11390_14
Total number of 3D structures: 78
LGA calculations using distance cutoff DIST: 4.0 A
Residues superimposed below 2.00 A: GREEN
Residues superimposed below 4.00 A: YELLOW
Residues superimposed below 6.00 A: ORANGE
Residues superimposed below 8.00 A: BROWN
Residues superimposed above 8.00 A or not aligned: RED
Terminal residues not aligned: GREY
Structure Deviation Summary
Calculations based on one final LGA superposition
(Bar representation of 3D plots, TEXT)
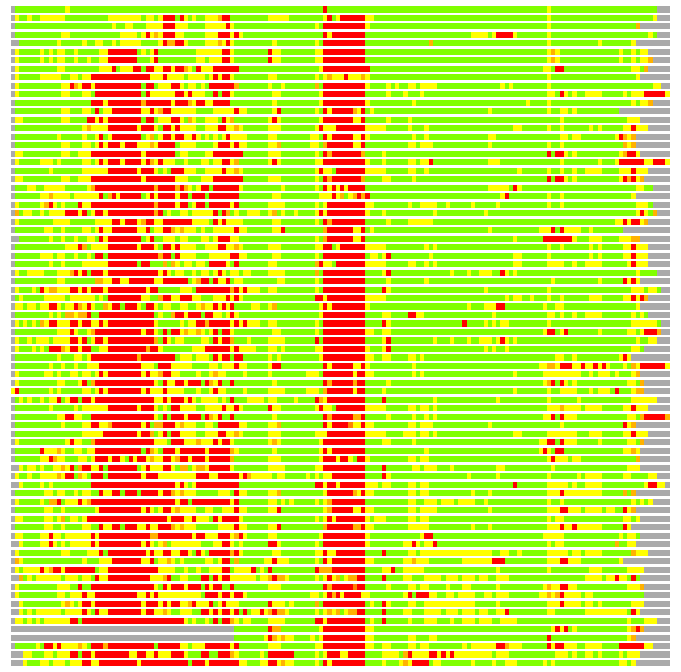
Structures ordered by LGA_S - score
| Structure |
NS |
NT |
N(dist=4.0) |
RMSD(N) |
Seq_ID(N) |
LGA_S |
LGA_Q |
PLOTS |
| 2cy2_A |
174 |
156 |
151 |
0.52 |
21.85 |
96.317 |
24.246 |
T P |
| 1tiq_A |
173 |
156 |
139 |
1.70 |
12.23 |
81.663 |
7.730 |
T P |
| 3f0a_A |
159 |
156 |
134 |
1.44 |
11.94 |
81.360 |
8.717 |
T P |
| 3fnc_B |
161 |
156 |
131 |
1.51 |
16.79 |
79.234 |
8.129 |
T P |
| 3fix_A |
165 |
156 |
132 |
1.72 |
11.36 |
78.320 |
7.266 |
T P |
| 1j4j_A |
170 |
156 |
129 |
1.57 |
14.73 |
77.769 |
7.726 |
T P |
| 1ghe_B |
171 |
156 |
130 |
1.65 |
14.62 |
77.421 |
7.437 |
T P |
| 2fiw_A |
160 |
156 |
121 |
1.67 |
24.79 |
72.910 |
6.851 |
T P |
| 1s3z_A |
147 |
156 |
129 |
1.92 |
16.28 |
70.558 |
6.375 |
T P |
| 2i6c_A |
160 |
156 |
119 |
1.74 |
17.65 |
70.018 |
6.460 |
T P |
| 2dxq_A |
147 |
156 |
121 |
1.85 |
15.70 |
69.106 |
6.203 |
T P |
| 2cnt_A |
151 |
156 |
115 |
1.68 |
13.91 |
68.936 |
6.479 |
T P |
| 2fxf_A |
164 |
156 |
123 |
1.88 |
14.63 |
65.204 |
6.223 |
T P |
| 1z4e_A |
150 |
156 |
123 |
1.78 |
17.07 |
65.180 |
6.544 |
T P |
| 2j8m_A |
171 |
156 |
126 |
1.94 |
17.46 |
65.091 |
6.187 |
T P |
| 2fe7_B |
166 |
156 |
125 |
2.00 |
18.40 |
64.987 |
5.959 |
T P |
| 2b3u_B |
164 |
156 |
125 |
1.98 |
13.60 |
64.946 |
6.017 |
T P |
| 3d2m_A |
424 |
156 |
109 |
1.75 |
18.35 |
64.663 |
5.896 |
T P |
| 2eui_A |
153 |
156 |
122 |
1.92 |
17.21 |
64.555 |
6.042 |
T P |
| 1yr0_A |
163 |
156 |
127 |
1.93 |
18.11 |
64.266 |
6.259 |
T P |
| 2r98_A |
424 |
156 |
110 |
1.85 |
18.18 |
64.243 |
5.631 |
T P |
| 2ob0_C |
166 |
156 |
116 |
1.83 |
10.34 |
64.163 |
6.008 |
T P |
| 1wwz_A |
157 |
156 |
119 |
1.89 |
19.33 |
64.111 |
5.972 |
T P |
| 1y9k_B |
154 |
156 |
111 |
1.90 |
17.12 |
64.093 |
5.546 |
T P |
| 1yre_B |
187 |
156 |
121 |
2.03 |
8.26 |
64.055 |
5.676 |
T P |
| 2q7b_A |
164 |
156 |
122 |
1.87 |
12.30 |
64.026 |
6.181 |
T P |
| 3bj7_A |
162 |
156 |
123 |
1.96 |
15.45 |
63.869 |
5.970 |
T P |
| 3fyn_A |
152 |
156 |
120 |
1.83 |
15.83 |
63.789 |
6.229 |
T P |
| 1yvo_B |
171 |
156 |
125 |
1.97 |
16.00 |
63.403 |
6.034 |
T P |
| 2ge3_A |
164 |
156 |
119 |
1.86 |
17.65 |
63.221 |
6.086 |
T P |
| 2jlm_D |
182 |
156 |
127 |
2.02 |
18.11 |
63.113 |
5.984 |
T P |
| 1s7f_A |
181 |
156 |
121 |
1.94 |
12.40 |
62.910 |
5.940 |
T P |
| 3d8p_B |
160 |
156 |
118 |
1.79 |
14.41 |
62.904 |
6.239 |
T P |
| 2z0z_A |
194 |
156 |
114 |
1.90 |
10.53 |
62.564 |
5.699 |
T P |
| 2i79_A |
171 |
156 |
122 |
2.00 |
13.11 |
62.310 |
5.808 |
T P |
| 3exn_A |
154 |
156 |
123 |
2.00 |
14.63 |
62.066 |
5.860 |
T P |
| 2vi7_C |
165 |
156 |
116 |
1.83 |
18.97 |
61.930 |
6.012 |
T P |
| 2z10_A |
187 |
156 |
114 |
1.92 |
9.65 |
61.797 |
5.652 |
T P |
| 2vez_A |
165 |
156 |
117 |
1.92 |
14.53 |
61.786 |
5.797 |
T P |
| 1z9u_B |
176 |
156 |
119 |
1.92 |
11.76 |
61.777 |
5.880 |
T P |
| 2zxv_A |
187 |
156 |
113 |
1.86 |
10.62 |
61.525 |
5.774 |
T P |
| 2pdo_D |
142 |
156 |
111 |
1.71 |
20.72 |
61.126 |
6.137 |
T P |
| 2bei_B |
159 |
156 |
120 |
2.00 |
16.67 |
61.036 |
5.710 |
T P |
| 1qso_A |
149 |
156 |
127 |
1.99 |
13.39 |
61.021 |
6.067 |
T P |
| 1i12_D |
157 |
156 |
115 |
1.93 |
15.65 |
61.016 |
5.656 |
T P |
| 1qsm_D |
152 |
156 |
128 |
1.97 |
12.50 |
60.992 |
6.182 |
T P |
| 1nsl_C |
180 |
156 |
116 |
1.95 |
12.07 |
60.931 |
5.662 |
T P |
| 2bl1_A |
170 |
156 |
126 |
2.09 |
17.46 |
60.269 |
5.763 |
T P |
| 1i1d_D |
161 |
156 |
114 |
2.00 |
15.79 |
60.219 |
5.431 |
T P |
| 2b5g_B |
159 |
156 |
122 |
2.03 |
13.93 |
59.819 |
5.738 |
T P |
| 3dr6_A |
172 |
156 |
125 |
2.09 |
15.20 |
59.549 |
5.712 |
T P |
| 3fb3_B |
149 |
156 |
115 |
1.88 |
20.00 |
59.299 |
5.796 |
T P |
| 1q2y_A |
140 |
156 |
109 |
1.82 |
17.43 |
59.101 |
5.687 |
T P |
| 1vkc_A |
149 |
156 |
119 |
2.02 |
16.81 |
58.965 |
5.620 |
T P |
| 3eo4_D |
162 |
156 |
117 |
2.09 |
11.11 |
58.923 |
5.345 |
T P |
| 3fbu_A |
166 |
156 |
118 |
2.12 |
9.32 |
58.920 |
5.322 |
T P |
| 3ec4_A |
224 |
156 |
109 |
1.80 |
13.76 |
58.656 |
5.731 |
T P |
| 2g3t_A |
169 |
156 |
126 |
2.17 |
13.49 |
58.572 |
5.562 |
T P |
| 1yvk_A |
152 |
156 |
109 |
1.90 |
12.84 |
58.542 |
5.445 |
T P |
| 2b5g_A |
164 |
156 |
123 |
2.13 |
14.63 |
58.288 |
5.522 |
T P |
| 2ae6_A |
148 |
156 |
112 |
2.03 |
16.07 |
57.980 |
5.265 |
T P |
| 2jev_A |
169 |
156 |
124 |
2.15 |
14.52 |
57.814 |
5.500 |
T P |
| 1mk4_A |
157 |
156 |
108 |
1.97 |
19.44 |
57.048 |
5.214 |
T P |
| 2r1i_A |
156 |
156 |
117 |
1.97 |
15.38 |
56.936 |
5.656 |
T P |
| 1vhs_A |
165 |
156 |
119 |
2.14 |
15.13 |
56.276 |
5.315 |
T P |
| 3dsb_A |
152 |
156 |
119 |
2.11 |
14.29 |
55.854 |
5.391 |
T P |
| 2fck_A |
174 |
156 |
113 |
2.09 |
10.62 |
55.289 |
5.153 |
T P |
| 2prb_A |
173 |
156 |
123 |
2.38 |
15.45 |
55.264 |
4.961 |
T P |
| 1i21_B |
157 |
156 |
112 |
2.00 |
16.07 |
54.819 |
5.343 |
T P |
| 1u6m_A |
189 |
156 |
118 |
2.26 |
19.49 |
54.504 |
5.005 |
T P |
| 3eg7_B |
175 |
156 |
113 |
2.17 |
13.27 |
53.601 |
4.969 |
T P |
| 2bue_A |
179 |
156 |
121 |
2.46 |
13.22 |
53.438 |
4.720 |
T P |
| 1s7k_A |
158 |
156 |
111 |
2.05 |
11.71 |
53.249 |
5.165 |
T P |
| 1y9w_A |
140 |
156 |
83 |
1.61 |
14.46 |
50.835 |
4.849 |
T P |
| 2g3a_A |
137 |
156 |
83 |
1.75 |
18.07 |
49.900 |
4.476 |
T P |
| 2ozh_A |
142 |
156 |
101 |
2.23 |
19.80 |
49.845 |
4.329 |
T P |
| 2pr8_A |
173 |
156 |
105 |
2.41 |
13.33 |
46.920 |
4.180 |
T P |
| 1yx0_A |
151 |
156 |
98 |
2.21 |
12.24 |
46.758 |
4.248 |
T P |
NS : Total number of residues in Structure (rotated structure)
NT : Total number of residues in TARGET (frame of reference)
N : Total number of residues superimposed under 4.0 Angstrom distance cutoff
RMSD : RMS deviation calculated on all N residues superimposed under 4.0 Angstrom distance cutoff
Seq_Id : Sequence Identity. Percent of identical residues from the total of N aligned.
LGA_S : Structure similarity score calculated by internal LGA procedure (see LGA paper for details)
LGA_Q : Score (how tight is the superposition) calculated by the formula: Q = 0.1*N/(0.1+RMSD)
PLOTS : T - Flat text file (output from LGA program, rotated structure)
PLOTS : P - Plot of superimposed structures (3D plot colored as bars)
Citing LGA:
Zemla A., "LGA - a Method for Finding 3D Similarities in Protein Structures",
Nucleic Acids Research, 2003, Vol. 31, No. 13, pp. 3370-3374.
[MEDLINE]

