LGA
Sequence Independent Analysis (LGA)
Frame of reference: Cat.Q2246_545_539.5wLII_11390_79
Total number of 3D structures: 50
LGA calculations using distance cutoff DIST: 4.0 A
Residues superimposed below 2.00 A: GREEN
Residues superimposed below 4.00 A: YELLOW
Residues superimposed below 6.00 A: ORANGE
Residues superimposed below 8.00 A: BROWN
Residues superimposed above 8.00 A or not aligned: RED
Terminal residues not aligned: GREY
Structure Deviation Summary
Calculations based on one final LGA superposition
(Bar representation of 3D plots, TEXT)
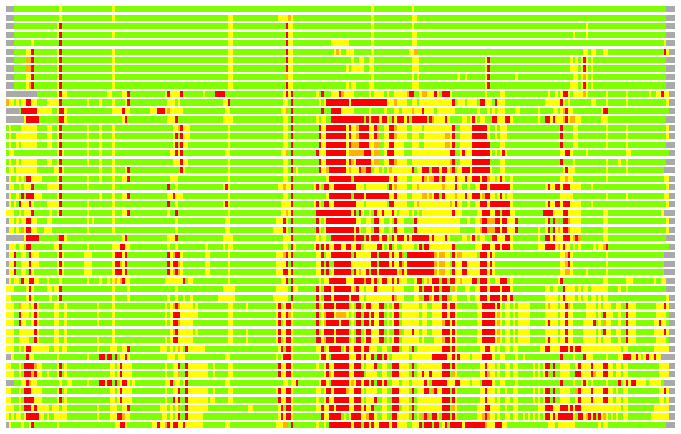
Structures ordered by LGA_S - score
| Structure |
NS |
NT |
N(dist=4.0) |
RMSD(N) |
Seq_ID(N) |
LGA_S |
LGA_Q |
PLOTS |
| 1zoi_A |
275 |
265 |
259 |
0.41 |
22.39 |
97.397 |
50.437 |
T P |
| 1a88_A |
275 |
265 |
259 |
0.88 |
23.55 |
95.977 |
26.459 |
T P |
| 1a8s_A |
273 |
265 |
257 |
0.86 |
21.40 |
95.480 |
26.696 |
T P |
| 1va4_A |
271 |
265 |
257 |
0.99 |
20.23 |
94.604 |
23.568 |
T P |
| 1a8q_A |
274 |
265 |
257 |
1.09 |
19.07 |
93.750 |
21.565 |
T P |
| 3fob_A |
277 |
265 |
256 |
1.26 |
17.97 |
92.483 |
18.783 |
T P |
| 1bro_A |
277 |
265 |
254 |
1.25 |
20.08 |
91.591 |
18.762 |
T P |
| 1a8u_A |
277 |
265 |
254 |
1.27 |
20.08 |
91.495 |
18.480 |
T P |
| 1hkh_A |
279 |
265 |
256 |
1.34 |
19.53 |
91.473 |
17.778 |
T P |
| 1brt_A |
277 |
265 |
254 |
1.27 |
20.08 |
91.455 |
18.504 |
T P |
| 1m33_A |
256 |
265 |
230 |
1.86 |
19.13 |
77.513 |
11.737 |
T P |
| 1j1i_A |
258 |
265 |
229 |
1.89 |
18.34 |
76.946 |
11.510 |
T P |
| 1wom_A |
271 |
265 |
236 |
1.89 |
17.37 |
74.556 |
11.847 |
T P |
| 3bf8_A |
255 |
265 |
204 |
1.74 |
19.12 |
71.514 |
11.109 |
T P |
| 2puj_A |
283 |
265 |
235 |
1.85 |
14.04 |
69.020 |
12.042 |
T P |
| 2rhw_A |
283 |
265 |
234 |
1.84 |
14.10 |
68.674 |
12.092 |
T P |
| 2pu6_A |
283 |
265 |
233 |
1.86 |
14.16 |
67.575 |
11.890 |
T P |
| 1u2e_A |
286 |
265 |
231 |
1.90 |
19.48 |
67.397 |
11.536 |
T P |
| 2og1_A |
285 |
265 |
232 |
1.83 |
14.66 |
67.284 |
12.007 |
T P |
| 2vf2_A |
284 |
265 |
225 |
1.81 |
16.89 |
67.184 |
11.784 |
T P |
| 1iup_A |
271 |
265 |
221 |
1.87 |
17.65 |
66.205 |
11.230 |
T P |
| 3cxu_A |
320 |
265 |
226 |
1.93 |
15.49 |
66.105 |
11.134 |
T P |
| 2d0d_A |
271 |
265 |
221 |
1.91 |
18.10 |
65.837 |
11.020 |
T P |
| 2cjp_A |
320 |
265 |
224 |
1.92 |
16.07 |
65.697 |
11.078 |
T P |
| 1ehy_A |
282 |
265 |
222 |
1.87 |
17.57 |
65.548 |
11.261 |
T P |
| 2e3j_A |
346 |
265 |
230 |
1.95 |
16.52 |
65.296 |
11.225 |
T P |
| 2zjf_A |
346 |
265 |
230 |
1.97 |
15.65 |
64.989 |
11.093 |
T P |
| 3bf7_A |
255 |
265 |
201 |
1.72 |
19.90 |
64.601 |
11.056 |
T P |
| 1y37_A |
294 |
265 |
225 |
1.91 |
18.22 |
63.841 |
11.203 |
T P |
| 2ocl_A |
254 |
265 |
213 |
1.91 |
18.31 |
63.670 |
10.580 |
T P |
| 2ocg_A |
254 |
265 |
213 |
1.90 |
18.78 |
63.605 |
10.671 |
T P |
| 2ock_A |
254 |
265 |
209 |
1.86 |
19.14 |
62.997 |
10.686 |
T P |
| 1c4x_A |
281 |
265 |
218 |
1.90 |
17.43 |
62.894 |
10.895 |
T P |
| 1zd3_A |
547 |
265 |
226 |
1.92 |
17.70 |
62.770 |
11.173 |
T P |
| 1cqz_B |
541 |
265 |
229 |
1.97 |
17.47 |
62.713 |
11.037 |
T P |
| 1bez_A |
310 |
265 |
227 |
2.08 |
14.54 |
62.188 |
10.399 |
T P |
| 2yxp_X |
310 |
265 |
231 |
2.17 |
13.42 |
61.586 |
10.198 |
T P |
| 1hde_A |
310 |
265 |
225 |
2.07 |
14.22 |
61.363 |
10.352 |
T P |
| 1be0_A |
310 |
265 |
226 |
2.11 |
14.16 |
61.267 |
10.237 |
T P |
| 1b6g_A |
308 |
265 |
228 |
2.12 |
14.04 |
61.165 |
10.286 |
T P |
| 1cqw_A |
295 |
265 |
226 |
2.15 |
18.58 |
60.414 |
10.024 |
T P |
| 1azw_A |
313 |
265 |
212 |
2.04 |
19.34 |
60.264 |
9.919 |
T P |
| 1cv2_A |
293 |
265 |
225 |
2.16 |
16.44 |
59.984 |
9.935 |
T P |
| 1mj5_A |
297 |
265 |
225 |
2.18 |
16.44 |
59.976 |
9.870 |
T P |
| 1wm1_A |
313 |
265 |
211 |
2.04 |
18.01 |
59.900 |
9.871 |
T P |
| 1g42_A |
293 |
265 |
223 |
2.16 |
17.04 |
59.701 |
9.878 |
T P |
| 1iz7_A |
294 |
265 |
224 |
2.16 |
16.52 |
59.664 |
9.921 |
T P |
| 1bn6_A |
291 |
265 |
223 |
2.20 |
17.94 |
59.245 |
9.711 |
T P |
| 2v9z_A |
302 |
265 |
214 |
2.11 |
17.76 |
58.841 |
9.693 |
T P |
| 2r11_D |
288 |
265 |
208 |
1.99 |
17.79 |
57.658 |
9.971 |
T P |
NS : Total number of residues in Structure (rotated structure)
NT : Total number of residues in TARGET (frame of reference)
N : Total number of residues superimposed under 4.0 Angstrom distance cutoff
RMSD : RMS deviation calculated on all N residues superimposed under 4.0 Angstrom distance cutoff
Seq_Id : Sequence Identity. Percent of identical residues from the total of N aligned.
LGA_S : Structure similarity score calculated by internal LGA procedure (see LGA paper for details)
LGA_Q : Score (how tight is the superposition) calculated by the formula: Q = 0.1*N/(0.1+RMSD)
PLOTS : T - Flat text file (output from LGA program, rotated structure)
PLOTS : P - Plot of superimposed structures (3D plot colored as bars)
Citing LGA:
Zemla A., "LGA - a Method for Finding 3D Similarities in Protein Structures",
Nucleic Acids Research, 2003, Vol. 31, No. 13, pp. 3370-3374.
[MEDLINE]

