LGA
Sequence Independent Analysis (LGA)
Frame of reference: Cat.Q2246_545_69.5wLII_10961_34
Total number of 3D structures: 25
LGA calculations using distance cutoff DIST: 4.0 A
Residues superimposed below 2.00 A: GREEN
Residues superimposed below 4.00 A: YELLOW
Residues superimposed below 6.00 A: ORANGE
Residues superimposed below 8.00 A: BROWN
Residues superimposed above 8.00 A or not aligned: RED
Terminal residues not aligned: GREY
Structure Deviation Summary
Calculations based on one final LGA superposition
(Bar representation of 3D plots, TEXT)
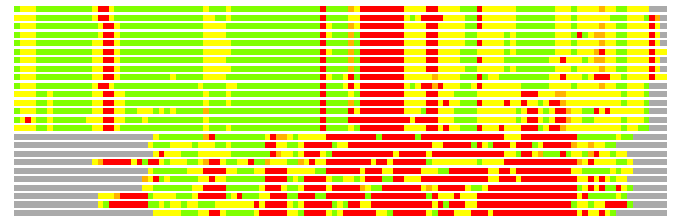
Structures ordered by LGA_S - score
| Structure |
NS |
NT |
N(dist=4.0) |
RMSD(N) |
Seq_ID(N) |
LGA_S |
LGA_Q |
PLOTS |
| 1q9h_A |
430 |
117 |
100 |
1.98 |
22.00 |
65.624 |
4.807 |
T P |
| 1gpi_A |
431 |
117 |
99 |
1.98 |
21.21 |
65.442 |
4.762 |
T P |
| 7cel_A |
434 |
117 |
101 |
2.11 |
19.80 |
64.504 |
4.568 |
T P |
| 4cel_A |
434 |
117 |
101 |
2.11 |
17.82 |
64.469 |
4.565 |
T P |
| 6cel_A |
434 |
117 |
101 |
2.10 |
19.80 |
64.240 |
4.588 |
T P |
| 2v3i_A |
434 |
117 |
102 |
2.12 |
18.63 |
64.147 |
4.601 |
T P |
| 1q2e_A |
426 |
117 |
100 |
2.10 |
19.00 |
63.995 |
4.539 |
T P |
| 1egn_A |
434 |
117 |
102 |
2.14 |
19.61 |
63.931 |
4.558 |
T P |
| 1q2b_A |
434 |
117 |
99 |
2.05 |
17.17 |
63.700 |
4.612 |
T P |
| 2rfw_A |
430 |
117 |
98 |
1.99 |
22.45 |
63.600 |
4.681 |
T P |
| 1ojj_A |
398 |
117 |
98 |
2.29 |
15.31 |
62.555 |
4.106 |
T P |
| 1dym_A |
398 |
117 |
95 |
2.20 |
14.74 |
61.061 |
4.133 |
T P |
| 3ovw_A |
400 |
117 |
94 |
2.11 |
12.77 |
60.735 |
4.248 |
T P |
| 1a39_A |
402 |
117 |
91 |
2.19 |
16.48 |
60.049 |
3.981 |
T P |
| 2a39_A |
398 |
117 |
93 |
2.15 |
13.98 |
59.840 |
4.137 |
T P |
| 2p4b_A |
284 |
117 |
44 |
2.03 |
9.09 |
29.950 |
2.066 |
T P |
| 2yzy_A |
163 |
117 |
45 |
2.43 |
4.44 |
28.935 |
1.775 |
T P |
| 2v43_B |
276 |
117 |
47 |
2.56 |
8.51 |
28.633 |
1.765 |
T P |
| 2byo_A |
183 |
117 |
55 |
2.85 |
10.91 |
28.555 |
1.865 |
T P |
| 2zf3_A |
186 |
117 |
46 |
2.45 |
6.52 |
27.332 |
1.806 |
T P |
| 3buu_A |
224 |
117 |
45 |
2.49 |
11.11 |
27.054 |
1.737 |
T P |
| 3bmz_B |
190 |
117 |
41 |
2.25 |
4.88 |
26.421 |
1.745 |
T P |
| 2zpd_A |
183 |
117 |
40 |
2.38 |
7.50 |
24.544 |
1.615 |
T P |
| 3bk5_A |
235 |
117 |
41 |
2.45 |
4.88 |
23.762 |
1.609 |
T P |
| 1iwl_A |
177 |
117 |
37 |
2.62 |
0.00 |
23.663 |
1.359 |
T P |
NS : Total number of residues in Structure (rotated structure)
NT : Total number of residues in TARGET (frame of reference)
N : Total number of residues superimposed under 4.0 Angstrom distance cutoff
RMSD : RMS deviation calculated on all N residues superimposed under 4.0 Angstrom distance cutoff
Seq_Id : Sequence Identity. Percent of identical residues from the total of N aligned.
LGA_S : Structure similarity score calculated by internal LGA procedure (see LGA paper for details)
LGA_Q : Score (how tight is the superposition) calculated by the formula: Q = 0.1*N/(0.1+RMSD)
PLOTS : T - Flat text file (output from LGA program, rotated structure)
PLOTS : P - Plot of superimposed structures (3D plot colored as bars)
Citing LGA:
Zemla A., "LGA - a Method for Finding 3D Similarities in Protein Structures",
Nucleic Acids Research, 2003, Vol. 31, No. 13, pp. 3370-3374.
[MEDLINE]

