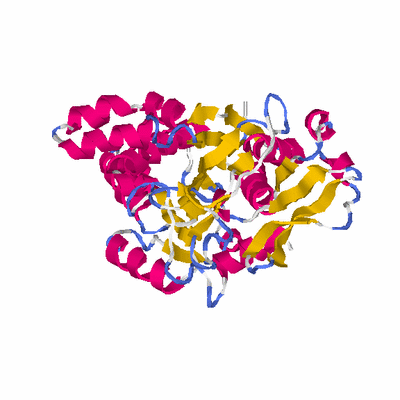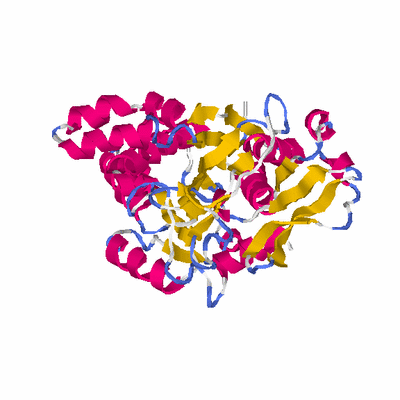##########
# PROTEIN: Q2246_545_130.5wLII_11077_46 359 130 # Category_B
# Length: 359
# Weight: 40567.50
# Similarity: UniProt_B6ARF1, BLAST_B6ARF1, PDB_templates_INFO
# Best Pcover Model PDB N_AA SISC E-val Seq_ID LAL Overlap
# Cat 88.02 M_00 1zkd_A 387 2 6e-94 28.000 316:347 (2-340:22-353)
# Evl 87.47 M_00 1zkd_A 387 2 e-105 28.000 314:349 (2-340:22-353)
# Sid 76.32 M_00 1zkd_A 387 2 7e-25 31.000 274:303 (6-305:26-310)
# Cov 88.02 M_00 1zkd_A 387 2 6e-94 28.000 316:347 (2-340:22-353)
# LaL 22.56 M_04 1i9g_A 280 1 0.030 15.000 81:88 (51-138:89-169)
# OvN 87.47 M_00 1zkd_A 387 2 e-105 28.000 314:349 (2-340:22-353)
# OvC 87.47 M_00 1zkd_A 387 2 e-105 28.000 314:349 (2-340:22-353)
PDB: 1zkd PDBsum
HEADER STRUCTURAL GENOMICS, UNKNOWN FUNCTION 02-MAY-05 1ZKD
TITLE X-RAY STRUCTURE OF THE PUTATIVE PROTEIN Q6N1P6 FROM
TITLE 2 RHODOPSEUDOMONAS PALUSTRIS AT THE RESOLUTION 2.1 A ,
TITLE 3 NORTHEAST STRUCTURAL GENOMICS CONSORTIUM TARGET RPR58
SOURCE 2 ORGANISM_SCIENTIFIC: RHODOPSEUDOMONAS PALUSTRIS CGA009;
SOURCE 3 ORGANISM_TAXID: 258594;
KEYWDS X-RAY, NESG, RPR58, STRUCTURAL GENOMICS, PSI, PROTEIN
KEYWDS 2 STRUCTURE INITIATIVE, NORTHEAST STRUCTURAL GENOMICS
KEYWDS 3 CONSORTIUM, UNKNOWN FUNCTION
SCOP: c.66 S-adenosyl-L-methionine-dependent methyltransferases
SCOP: c.66.1 S-adenosyl-L-methionine-dependent methyltransferases
SCOP: c.66.1.52 RPA4359-like
PDB: 1i9g PDBsum
HEADER TRANSFERASE 20-MAR-01 1I9G
TITLE CRYSTAL STRUCTURE OF AN ADOMET DEPENDENT METHYLTRANSFERASE
SOURCE 2 ORGANISM_SCIENTIFIC: MYCOBACTERIUM TUBERCULOSIS;
SOURCE 3 ORGANISM_TAXID: 1773;
KEYWDS MYCOBACTERIUM, MTASE, ADOMET, CRYSTAL, STRUCTURAL GENOMICS,
KEYWDS 2 PSI, PROTEIN STRUCTURE INITIATIVE, TB STRUCTURAL GENOMICS
KEYWDS 3 CONSORTIUM, TBSGC, TRANSFERASE
SCOP: c.66 S-adenosyl-L-methionine-dependent methyltransferases
SCOP: c.66.1 S-adenosyl-L-methionine-dependent methyltransferases
SCOP: c.66.1.13 tRNA(1-methyladenosine) methyltransferase-like