##########
# PROTEIN: Q2246_545_145.5wLII_11111_49 140 145 # Category_C1
# Length: 140
# Weight: 15112.96
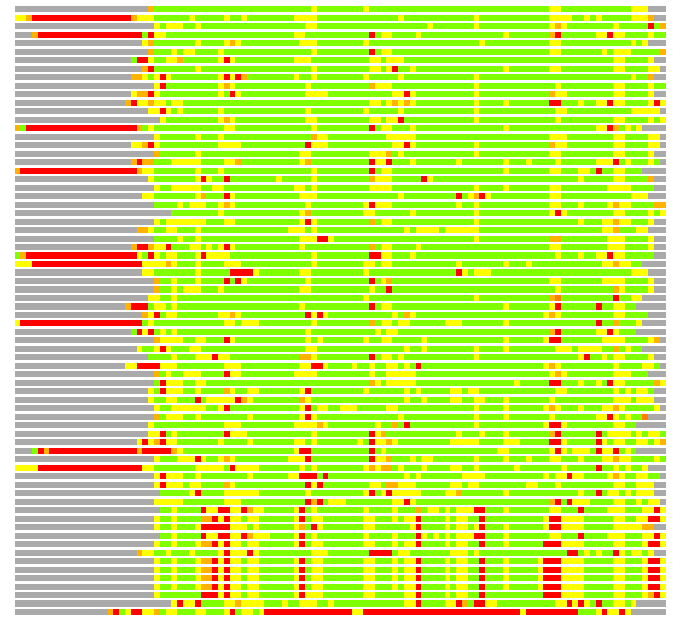
# Similarity: UniProt_B6AKE2, BLAST_B6AKE2, PDB_templates_INFO
# Best Pcover Model PDB N_AA SISC E-val Seq_ID LAL Overlap
# Cat 75.71 M_00 2nzi_A 305 2 2e-16 16.000 106:119 (30-136:176-294)
# Evl 67.14 M_00 1x6l_A 563 3 2e-20 12.000 94:127 (29-136:10-136)
# Sid 48.57 M_01 z10w 419 0 0.079 32.000 68:77 (27-97:334-408)
# Cov 90.00 M_18 3fl7_A 536 1 0.005 11.000 126:145 (2-136:394-534)
# LaL 32.14 M_36 1wfu_A 120 1 0.083 20.000 45:45 (90-134:68-112)
# OvN 87.14 M_03 3fl7_A 536 1 5e-15 10.000 122:149 (2-136:394-534)
# OvC 82.86 M_06 1bqu_B 215 2 1e-13 12.000 116:138 (25-140:73-210)
PDB: 2nzi PDBsum
HEADER TRANSFERASE 23-NOV-06 2NZI
TITLE CRYSTAL STRUCTURE OF DOMAINS A168-A170 FROM TITIN
SOURCE 2 ORGANISM_SCIENTIFIC: HOMO SAPIENS;
SOURCE 3 ORGANISM_COMMON: HUMAN;
SOURCE 4 ORGANISM_TAXID: 9606;
COMPND 6 EC: 2.7.11.1;
KEYWDS IG-DOMAIN, FNIII-DOMAIN, TRANSFERASE
SCOP: none
PDB: 1x6l PDBsum
HEADER HYDROLASE 11-AUG-04 1X6L
TITLE CRYSTAL STRUCTURE OF S. MARCESCENS CHITINASE A MUTANT W167A
SOURCE 2 ORGANISM_SCIENTIFIC: SERRATIA MARCESCENS;
SOURCE 3 ORGANISM_TAXID: 615;
COMPND 4 EC: 3.2.1.14;
KEYWDS CHITINASE A, HYDROLASE
SCOP: b.1 Immunoglobulin-like beta-sandwich
SCOP: b.1.18 E set domains
SCOP: b.1.18.2 E-set domains of sugar-utilizing enzymes
SCOP: c.1 TIM beta/alpha-barrel
SCOP: c.1.8 (Trans)glycosidases
SCOP: c.1.8.5 Type II chitinase
SCOP: d.26 FKBP-like
SCOP: d.26.3 Chitinase insertion domain
SCOP: d.26.3.1 Chitinase insertion domain
PDB: 1cz1 PDBsum
HEADER HYDROLASE 01-SEP-99 1CZ1
TITLE EXO-B-(1,3)-GLUCANASE FROM CANDIDA ALBICANS AT 1.85 A
TITLE 2 RESOLUTION
SOURCE 2 ORGANISM_SCIENTIFIC: CANDIDA ALBICANS;
SOURCE 3 ORGANISM_TAXID: 5476;
COMPND 4 EC: 3.2.1.58;
KEYWDS EXOGLUCANASE, CANDIDA ALBICANS, GLYCOSIDE HYDROLASE FAMILY 5
SCOP: c.1 TIM beta/alpha-barrel
SCOP: c.1.8 (Trans)glycosidases
SCOP: c.1.8.3 beta-glycanases
PDB: 3fl7 PDBsum
HEADER TRANSFERASE, SIGNALING PROTEIN 18-DEC-08 3FL7
TITLE CRYSTAL STRUCTURE OF THE HUMAN EPHRIN A2 ECTODOMAIN
SOURCE 2 ORGANISM_SCIENTIFIC: HOMO SAPIENS;
SOURCE 3 ORGANISM_COMMON: HUMAN;
SOURCE 4 ORGANISM_TAXID: 9606;
COMPND 5 EC: 2.7.10.1;
KEYWDS ATP-BINDING, KINASE, NUCLEOTIDE-BINDING, RECEPTOR, SIGNAL,
KEYWDS 2 TRANSFERASE, PHOSPHORYLATION, TRANSMEMBRANE, TYROSINE-
KEYWDS 3 PROTEIN KINASE, GLYCOPROTEIN, LIGAND BINDING DOMAIN,
KEYWDS 4 CYSTEINE-RICH DOMAIN, SUSHI DOMAIN, EGF-LIKE MOTIF,
KEYWDS 5 FIBRONECTIN DOMAIN, STRUCTURAL GENOMICS CONSORTIUM, SGC,
KEYWDS 6 MEMBRANE, PHOSPHOPROTEIN, SIGNALING PROTEIN
SCOP: none
PDB: 1wfu PDBsum
HEADER STRUCTURAL GENOMICS, UNKNOWN FUNCTION 27-MAY-04 1WFU
TITLE SOLUTION STRUCTURE OF FIBRONECTIN TYPE III DOMAIN OF MOUSE
TITLE 2 HYPOTHETICAL PROTEIN
SOURCE 2 ORGANISM_SCIENTIFIC: MUS MUSCULUS;
SOURCE 3 ORGANISM_COMMON: HOUSE MOUSE;
SOURCE 4 ORGANISM_TAXID: 10090;
KEYWDS FN3 DOMAIN, SIMILAR TO 1700007B22RIK PROTEIN, STRUCTURAL
KEYWDS 2 GENOMICS, RIKEN STRUCTURAL GENOMICS/PROTEOMICS INITIATIVE,
KEYWDS 3 RSGI, UNKNOWN FUNCTION
SCOP: b.1 Immunoglobulin-like beta-sandwich
SCOP: b.1.2 Fibronectin type III
SCOP: b.1.2.1 Fibronectin type III
PDB: 1bqu PDBsum
HEADER SIGNALING PROTEIN 18-AUG-98 1BQU
TITLE CYTOKYNE-BINDING REGION OF GP130
SOURCE 2 ORGANISM_SCIENTIFIC: HOMO SAPIENS;
SOURCE 3 ORGANISM_COMMON: HUMAN;
SOURCE 4 ORGANISM_TAXID: 9606;
KEYWDS CYTOKINE RECEPTOR, GLYCOPROTEIN 130, GP130, INTERLEUKINE 6
KEYWDS 2 RECEPTOR BETA SUBUNIT, SIGNALING PROTEIN
SCOP: b.1 Immunoglobulin-like beta-sandwich
SCOP: b.1.2 Fibronectin type III
SCOP: b.1.2.1 Fibronectin type III