##########
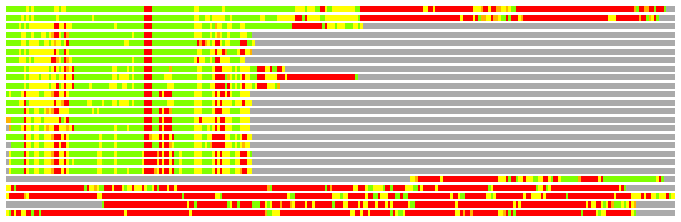
# PROTEIN: Q2246_545_191.5wLII_11181_78 268 191 # Category_C1
# Length: 268
# Weight: 29263.31
# Similarity: UniProt_A3EQ95, BLAST_A3EQ95, PDB_templates_INFO
# Best Pcover Model PDB N_AA SISC E-val Seq_ID LAL Overlap
# Cat 96.64 M_00 3fig_B 646 2 7e-48 15.000 259:398 (1-263:253-646)
# Evl 94.03 M_00 3fig_B 646 2 6e-58 14.000 252:416 (2-263:234-646)
# Sid 24.63 M_00 1sr9_B 644 2 1e-05 36.000 66:74 (60-125:312-385)
# Cov 96.64 M_00 3fig_B 646 2 7e-48 15.000 259:398 (1-263:253-646)
# LaL 24.63 M_11 2qf7_A 1165 2 6e-05 4.000 66:66 (9-74:737-802)
# OvN 96.64 M_00 3fig_B 646 2 7e-48 15.000 259:398 (1-263:253-646)
# OvC 26.12 M_15 1t4o_B 117 2 0.047 19.000 70:81 (198-268:8-87)
PDB: 3fig PDBsum
HEADER TRANSFERASE 11-DEC-08 3FIG
TITLE CRYSTAL STRUCTURE OF LEUCINE-BOUND LEUA FROM MYCOBACTERIUM
TITLE 2 TUBERCULOSIS
SOURCE 2 ORGANISM_SCIENTIFIC: MYCOBACTERIUM TUBERCULOSIS;
SOURCE 3 ORGANISM_TAXID: 1773;
COMPND 6 EC: 2.3.3.13;
KEYWDS TIM BARREL, AMINO-ACID BIOSYNTHESIS, BRANCHED-CHAIN AMINO
KEYWDS 2 ACID BIOSYNTHESIS, LEUCINE BIOSYNTHESIS, TRANSFERASE
SCOP: none
PDB: 1sr9 PDBsum
HEADER TRANSFERASE 22-MAR-04 1SR9
TITLE CRYSTAL STRUCTURE OF LEUA FROM MYCOBACTERIUM TUBERCULOSIS
SOURCE 2 ORGANISM_SCIENTIFIC: MYCOBACTERIUM TUBERCULOSIS;
SOURCE 3 ORGANISM_TAXID: 1773;
COMPND 6 EC: 2.3.3.13;
KEYWDS TIM BARREL, TRANSFERASE
SCOP: a.5 RuvA C-terminal domain-like
SCOP: a.5.7 post-HMGL domain-like
SCOP: a.5.7.1 DmpG/LeuA communication domain-like
SCOP: c.1 TIM beta/alpha-barrel
SCOP: c.1.10 Aldolase
SCOP: c.1.10.5 HMGL-like
SCOP: d.270 2-isopropylmalate synthase LeuA, allosteric (dimerisation) domain
SCOP: d.270.1 2-isopropylmalate synthase LeuA, allosteric (dimerisation) domain
SCOP: d.270.1.1 2-isopropylmalate synthase LeuA, allosteric (dimerisation) domain
PDB: 2qf7 PDBsum
HEADER LIGASE 27-JUN-07 2QF7
TITLE CRYSTAL STRUCTURE OF A COMPLETE MULTIFUNCTIONAL PYRUVATE
TITLE 2 CARBOXYLASE FROM RHIZOBIUM ETLI
SOURCE 2 ORGANISM_SCIENTIFIC: RHIZOBIUM ETLI CFN 42;
SOURCE 3 ORGANISM_TAXID: 347834;
COMPND 4 EC: 6.4.1.1;
KEYWDS MULTI-DOMAIN, MULTI-FUNCTIONAL, BIOTIN-DEPENDENT, LIGASE
SCOP: none
PDB: 1t4o PDBsum
HEADER HYDROLASE 30-APR-04 1T4O
TITLE CRYSTAL STRUCTURE OF RNT1P DSRBD
SOURCE 2 ORGANISM_SCIENTIFIC: SACCHAROMYCES CEREVISIAE;
SOURCE 3 ORGANISM_COMMON: BAKER'S YEAST;
SOURCE 4 ORGANISM_TAXID: 4932;
COMPND 6 EC: 3.1.26.3;
KEYWDS RNT1P, DSRBD, RNA-BINDING, HYDROLASE
SCOP: d.50 dsRBD-like
SCOP: d.50.1 dsRNA-binding domain-like
SCOP: d.50.1.1 Double-stranded RNA-binding domain (dsRBD)