##########
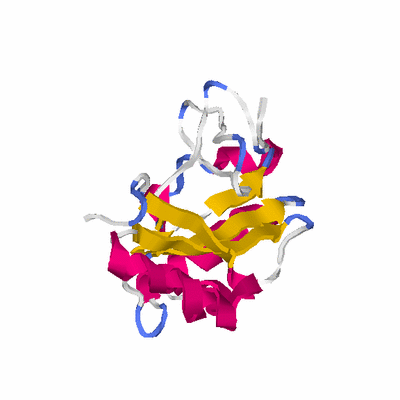
# PROTEIN: Q2246_545_240.5wLII_11212_56 220 240 # Category_C1
# Length: 220
# Weight: 22713.82
# Similarity: UniProt_B6APM0, BLAST_B6APM0, PDB_templates_INFO
# Best Pcover Model PDB N_AA SISC E-val Seq_ID LAL Overlap
# Cat 55.45 M_01 2yr1_A 257 2 6e-23 16.000 122:149 (42-185:111-237)
# Evl 75.00 M_00 2vjp_A 428 2 6e-41 12.000 165:196 (42-213:19-207)
# Sid 17.27 M_00 2gpt_A 523 1 0.051 38.000 38:39 (46-83:93-131)
# Cov 75.45 M_16 2g04_A 359 6 6e-34 12.000 166:194 (41-219:20-202)
# LaL 17.27 M_00 2gpt_A 523 1 0.051 38.000 38:39 (46-83:93-131)
# OvN 57.27 M_00 1gqn_A 252 7 1e-30 12.000 126:155 (38-185:102-234)
# OvC 73.18 M_14 2gci_A 360 24 2e-35 14.000 161:196 (41-219:18-200)
PDB: 2yr1 PDBsum
HEADER LYASE 01-APR-07 2YR1
TITLE CRYSTAL STRUCTURE OF 3-DEHYDROQUINATE DEHYDRATASE FROM
TITLE 2 GEOBACILLUS KAUSTOPHILUS HTA426
SOURCE 2 ORGANISM_SCIENTIFIC: GEOBACILLUS KAUSTOPHILUS HTA426;
SOURCE 3 ORGANISM_TAXID: 235909;
COMPND 5 EC: 4.2.1.10;
KEYWDS AMINO ACID BIOSYNTHESIS, 3-DEHYDROQUINASE, STRUCTURAL
KEYWDS 2 GENOMICS, NPPSFA, NATIONAL PROJECT ON PROTEIN STRUCTURAL
KEYWDS 3 AND FUNCTIONAL ANALYSES, RIKEN STRUCTURAL
KEYWDS 4 GENOMICS/PROTEOMICS INITIATIVE, RSGI, LYASE
SCOP: none
PDB: 2vjp PDBsum
HEADER TRANSFERASE 11-DEC-07 2VJP
TITLE FORMYL-COA TRANSFERASE MUTANT VARIANT W48F
SOURCE 2 ORGANISM_SCIENTIFIC: OXALOBACTER FORMIGENES;
SOURCE 3 ORGANISM_TAXID: 847;
COMPND 5 EC: 2.8.3.16;
KEYWDS CYTOPLASM, TRANSFERASE, CLASS III COA TRANSFERASE
SCOP: none
PDB: 2gpt PDBsum
HEADER OXIDOREDUCTASE, LYASE 18-APR-06 2GPT
TITLE CRYSTAL STRUCTURE OF ARABIDOPSIS DEHYDROQUINATE DEHYDRATASE-
TITLE 2 SHIKIMATE DEHYDROGENASE IN COMPLEX WITH TARTRATE AND
TITLE 3 SHIKIMATE
SOURCE 2 ORGANISM_SCIENTIFIC: ARABIDOPSIS THALIANA;
SOURCE 3 ORGANISM_COMMON: THALE CRESS;
SOURCE 4 ORGANISM_TAXID: 3702;
COMPND 5 EC: 4.2.1.10, 1.1.1.25;
KEYWDS TYPE I DEHYDROQUINATE DEHYDRATASE, AROE, SHIKIMATE
KEYWDS 2 DEHYDROGENASE, ARABIDOPSIS THALIANA, SHIKIMATE, TARTRATE,
KEYWDS 3 BIFUNCTIONAL ENZYME, OXIDOREDUCTASE, LYASE
SCOP: none
PDB: 2g04 PDBsum
HEADER ISOMERASE 11-FEB-06 2G04
TITLE CRYSTAL STRUCTURE OF FATTY ACID-COA RACEMASE FROM
TITLE 2 MYCOBACTERIUM TUBERCULOSIS H37RV
SOURCE 2 ORGANISM_SCIENTIFIC: MYCOBACTERIUM TUBERCULOSIS H37RV;
SOURCE 3 ORGANISM_TAXID: 83332;
COMPND 4 EC: 5.1.99.4;
KEYWDS ISOMERASE
SCOP: none
PDB: 1gqn PDBsum
HEADER LYASE 26-NOV-01 1GQN
TITLE NATIVE 3-DEHYDROQUINASE FROM SALMONELLA TYPHI
SOURCE 2 ORGANISM_SCIENTIFIC: SALMONELLA TYPHI;
SOURCE 3 ORGANISM_TAXID: 601;
COMPND 5 EC: 4.2.1.10;
KEYWDS LYASE, 3-DEHYDROQUINATE DEHYDRATASE, SHIKIMATE PATHWAY, A/B
KEYWDS 2 BARREL
SCOP: c.1 TIM beta/alpha-barrel
SCOP: c.1.10 Aldolase
SCOP: c.1.10.1 Class I aldolase
PDB: 2gci PDBsum
HEADER ISOMERASE 14-MAR-06 2GCI
TITLE THE 1,1-PROTON TRANSFER REACTION MECHANISM BY ALPHA-
TITLE 2 METHYLACYL-COA RACEMASE IS CATALYZED BY AN
TITLE 3 ASPARTE/HISTIDINE PAIR AND INVOLVES A SMOOTH, METHIONINE-
TITLE 4 RICH SURFACE FOR BINDING THE FATTY ACYL MOIETY
SOURCE 2 ORGANISM_SCIENTIFIC: MYCOBACTERIUM TUBERCULOSIS;
SOURCE 3 ORGANISM_TAXID: 1773;
COMPND 6 EC: 5.1.99.4;
KEYWDS ALPHA-METHYLACYL-COA RACEMASE, RACEMASE, COA TRANSFERASE,
KEYWDS 2 PROTON TRANSFER, COENZYME A, ISOMERASE
SCOP: c.123 CoA-transferase family III (CaiB/BaiF)
SCOP: c.123.1 CoA-transferase family III (CaiB/BaiF)
SCOP: c.123.1.1 CoA-transferase family III (CaiB/BaiF)