##########
# PROTEIN: Q2246_545_252.5wLII_11212_88 293 252 # Category_C1
# Length: 293
# Weight: 33783.51
# Similarity: UniProt_B6APQ1, BLAST_B6APQ1, PDB_templates_INFO
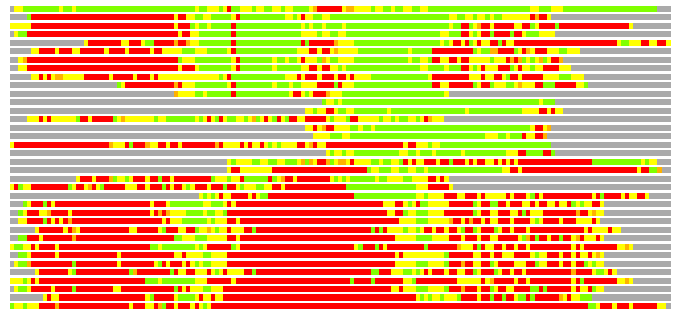
# Best Pcover Model PDB N_AA SISC E-val Seq_ID LAL Overlap
# Cat 52.56 M_00 1qvr_A 854 3 3e-05 15.000 154:161 (132-286:353-512)
# Evl 33.45 M_00 2e7s_D 135 20 4e-07 15.000 98:112 (187-287:6-114)
# Sid 40.61 M_08 2cd8_B 436 9 0.023 29.000 119:158 (17-155:97-234)
# Cov 70.99 M_22 1f5n_A 592 2 0.094 12.000 208:239 (63-286:361-584)
# LaL 35.84 M_06 1y1u_A 585 3 0.020 14.000 105:107 (170-274:11-117)
# OvN 40.61 M_08 2cd8_B 436 9 0.023 29.000 119:158 (17-155:97-234)
# OvC 49.49 M_05 2iw5_A 666 3 0.005 12.000 145:170 (146-293:176-342)
PDB: 1qvr PDBsum
HEADER CHAPERONE 28-AUG-03 1QVR
TITLE CRYSTAL STRUCTURE ANALYSIS OF CLPB
SOURCE 2 ORGANISM_SCIENTIFIC: THERMUS THERMOPHILUS;
SOURCE 3 ORGANISM_TAXID: 274;
KEYWDS COILED COIL, AAA ATPASE, CHAPERONE
SCOP: a.174 Double Clp-N motif
SCOP: a.174.1 Double Clp-N motif
SCOP: a.174.1.1 Double Clp-N motif
SCOP: c.37 P-loop containing nucleoside triphosphate hydrolases
SCOP: c.37.1 P-loop containing nucleoside triphosphate hydrolases
SCOP: c.37.1.20 Extended AAA-ATPase domain
PDB: 2e7s PDBsum
HEADER ENDOCYTOSIS/EXOCYTOSIS 12-JAN-07 2E7S
TITLE CRYSTAL STRUCTURE OF THE YEAST SEC2P GEF DOMAIN
SOURCE 2 ORGANISM_SCIENTIFIC: SACCHAROMYCES CEREVISIAE;
SOURCE 3 ORGANISM_COMMON: BAKER'S YEAST;
SOURCE 4 ORGANISM_TAXID: 4932;
KEYWDS COILED COIL, ENDOCYTOSIS/EXOCYTOSIS COMPLEX
SCOP: h.1 Parallel coiled-coil
SCOP: h.1.33 Sec2 N-terminal region
SCOP: h.1.33.1 Sec2 N-terminal region
PDB: 2cd8 PDBsum
HEADER OXIDOREDUCTASE 20-JAN-06 2CD8
TITLE CRYSTAL STRUCTURE OF YC-17-BOUND CYTOCHROME P450 PIKC
TITLE 2 (CYP107L1)
SOURCE 2 ORGANISM_SCIENTIFIC: STREPTOMYCES VENEZUELAE;
SOURCE 3 ORGANISM_TAXID: 54571;
KEYWDS OXIDOREDUCTASE, CYTOCHROME P450, PIKC, YC-17, MACROLIDE
KEYWDS 2 MONOOXYGENASE, ANTIBIOTIC BIOSYNTHESIS, HEME, IRON,
KEYWDS 3 METAL-BINDING
SCOP: none
PDB: 1f5n PDBsum
HEADER SIGNALING PROTEIN 15-JUN-00 1F5N
TITLE HUMAN GUANYLATE BINDING PROTEIN-1 IN COMPLEX WITH THE GTP
TITLE 2 ANALOGUE, GMPPNP.
SOURCE 2 ORGANISM_SCIENTIFIC: HOMO SAPIENS;
SOURCE 3 ORGANISM_COMMON: HUMAN;
SOURCE 4 ORGANISM_TAXID: 9606;
KEYWDS GBP, GTP HYDROLYSIS, GDP, GMP, INTERFERON INDUCED, DYNAMIN
KEYWDS 2 RELATED, LARGE GTPASE FAMILY. GMPPNP, GPPNHP., SIGNALING
KEYWDS 3 PROTEIN
SCOP: a.114 Interferon-induced guanylate-binding protein 1 (GBP1), C-terminal domain
SCOP: a.114.1 Interferon-induced guanylate-binding protein 1 (GBP1), C-terminal domain
SCOP: a.114.1.1 Interferon-induced guanylate-binding protein 1 (GBP1), C-terminal domain
SCOP: c.37 P-loop containing nucleoside triphosphate hydrolases
SCOP: c.37.1 P-loop containing nucleoside triphosphate hydrolases
SCOP: c.37.1.8 G proteins
PDB: 1y1u PDBsum
HEADER SIGNALING PROTEIN 19-NOV-04 1Y1U
TITLE STRUCTURE OF UNPHOSPHORYLATED STAT5A
SOURCE 2 ORGANISM_SCIENTIFIC: MUS MUSCULUS;
SOURCE 3 ORGANISM_COMMON: HOUSE MOUSE;
SOURCE 4 ORGANISM_TAXID: 10090;
KEYWDS ACTIVATOR, STAT, DNA-BINDING, SH2 DOMAIN, TRANSCRIPTION
KEYWDS 2 REGULATION, SIGNALING PROTEIN
SCOP: none
PDB: 2iw5 PDBsum
HEADER OXIDOREDUCTASE/TRANSCRIPTION REGULATOR 26-JUN-06 2IW5
TITLE STRUCTURAL BASIS FOR COREST-DEPENDENT DEMETHYLATION OF
TITLE 2 NUCLEOSOMES BY THE HUMAN LSD1 HISTONE DEMETHYLASE
SOURCE 2 ORGANISM_SCIENTIFIC: HOMO SAPIENS;
SOURCE 3 ORGANISM_COMMON: HUMAN;
SOURCE 4 ORGANISM_TAXID: 9606;
COMPND 8 EC: 1.-.-.-;
KEYWDS OXIDOREDUCTASE/TRANSCRIPTION REGULATOR,
KEYWDS 2 OXIDOREDUCTASE/REPRESSOR COMPLEX, HISTONE DEMETHYLASE,
KEYWDS 3 ALTERNATIVE SPLICING, FAD, LSD1, COREST, REPRESSOR, COILED
KEYWDS 4 COIL, TRANSCRIPTION REGULATION, HOST-VIRUS INTERACTION,
KEYWDS 5 CHROMATIN DEMETHYLATION, NUCLEAR PROTEIN, PHOSPHORYLATION,
KEYWDS 6 CHROMATIN REGULATOR, NUCLEOSOMES, TRANSCRIPTION,
KEYWDS 7 OXIDOREDUCTASE
SCOP: a.4 DNA/RNA-binding 3-helical bundle
SCOP: a.4.1 Homeodomain-like
SCOP: a.4.1.3 Myb/SANT domain