##########
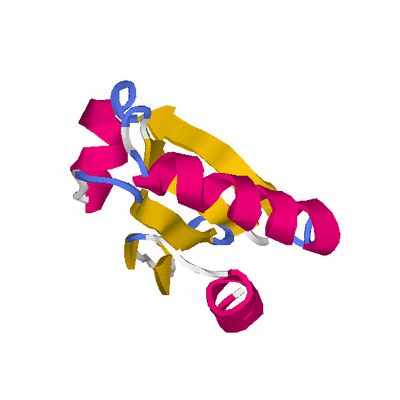
# PROTEIN: Q2246_545_283.5wLII_11238_16 83 283 # Category_B
# Length: 83
# Weight: 9512.98
# Similarity: UniProt_B6AQZ8, BLAST_B6AQZ8, PDB_templates_INFO
# Best Pcover Model PDB N_AA SISC E-val Seq_ID LAL Overlap
# Cat 96.39 M_28 2gzy_A 104 2 2e-32 21.000 80:80 (3-82:20-99)
# Evl 95.18 M_00 3dxb_D 222 8 7e-35 15.000 79:83 (3-81:33-115)
# Sid 90.36 M_33 2gzy_A 104 2 2e-28 22.000 75:79 (7-81:24-102)
# Cov 98.80 M_17 z0es 108 0 2e-31 10.000 82:82 (1-82:22-103)
# LaL 98.80 M_17 z0es 108 0 2e-31 10.000 82:82 (1-82:22-103)
# OvN 97.59 M_14 z0es 108 0 3e-34 14.000 81:85 (1-81:22-106)
# OvC 97.59 M_28 2i4a_A 107 1 4e-31 14.000 81:85 (3-83:23-107)
PDB: 2gzy PDBsum
HEADER ELECTRON TRANSPORT 12-MAY-06 2GZY
TITLE SOLUTION STRUCTURES OF THE REDUCED FORM OF THIOREDOXIN FROM
TITLE 2 BACILLUS SUBTILIS
SOURCE 2 ORGANISM_SCIENTIFIC: BACILLUS SUBTILIS;
SOURCE 3 ORGANISM_TAXID: 1423;
KEYWDS ALPHA/BETA, ELECTRON TRANSPORT
SCOP: none
PDB: 3dxb PDBsum
HEADER SPLICING, TRANSCRIPTION 24-JUL-08 3DXB
TITLE STRUCTURE OF THE UHM DOMAIN OF PUF60 FUSED TO THIOREDOXIN
SOURCE 2 ORGANISM_SCIENTIFIC: ESCHERICHIA COLI O157:H7;
SOURCE 3 ORGANISM_TAXID: 83334;
KEYWDS SPLICING, ALTERNATIVE SPLICING, FBP INTERACTING REPRESSOR,
KEYWDS 2 UHM, RRM, ELECTRON TRANSPORT, REDOX-ACTIVE CENTER,
KEYWDS 3 TRANSPORT, TRANSCRIPTION
SCOP: none
PDB: 2trx PDBsum
HEADER ELECTRON TRANSPORT 19-MAR-90 2TRX
TITLE CRYSTAL STRUCTURE OF THIOREDOXIN FROM ESCHERICHIA COLI AT
TITLE 2 1.68 ANGSTROMS RESOLUTION
SOURCE 2 ORGANISM_SCIENTIFIC: ESCHERICHIA COLI;
SOURCE 3 ORGANISM_TAXID: 562
KEYWDS ELECTRON TRANSPORT
SCOP: c.47 Thioredoxin fold
SCOP: c.47.1 Thioredoxin-like
SCOP: c.47.1.1 Thioltransferase
PDB: 2i4a PDBsum
HEADER OXIDOREDUCTASE 21-AUG-06 2I4A
TITLE CRYSTAL STRUCTURE OF THIOREDOXIN FROM THE ACIDOPHILE
TITLE 2 ACETOBACTER ACETI
SOURCE 2 ORGANISM_SCIENTIFIC: ACETOBACTER ACETI;
SOURCE 3 ORGANISM_TAXID: 435;
KEYWDS THIOREDOXIN, ACIDOPHILE, DISULFIDE EXCHANGE, OXIDOREDUCTASE
SCOP: none