##########
# PROTEIN: Q2246_545_3.5wLII_08492_1 366 3 # Category_C2
# Length: 366
# Weight: 42812.52
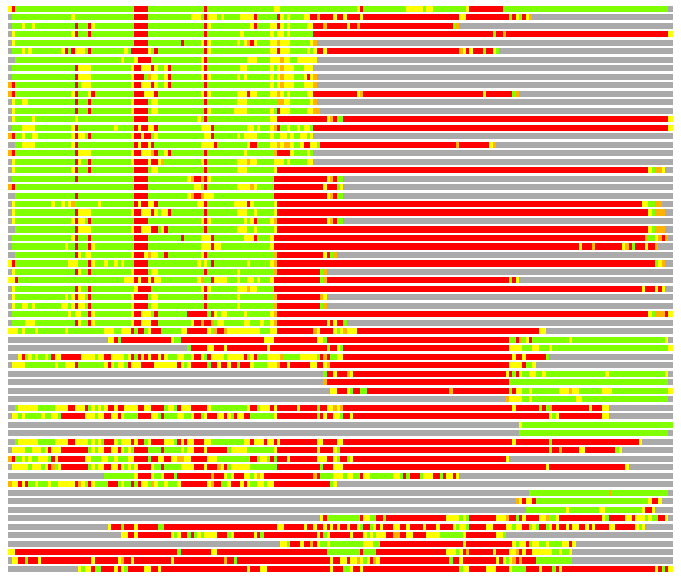
# Similarity: UniProt_B6ASI1, BLAST_B6ASI1, PDB_templates_INFO
# Best Pcover Model PDB N_AA SISC E-val Seq_ID LAL Overlap
# Cat 48.91 M_00 3c3w_A 225 2 6e-37 15.000 179:200 (170-364:9-192)
# Evl 46.99 M_00 z0ci 209 0 1e-51 10.000 172:204 (173-364:6-191)
# Sid 10.93 M_12 1s7o_C 113 3 0.012 32.000 40:40 (322-361:24-63)
# Cov 53.83 M_12 3clo_A 258 3 0.029 15.000 197:237 (122-358:38-234)
# LaL 14.21 M_10 3clo_A 258 3 1e-14 23.000 52:52 (313-364:189-240)
# OvN 53.83 M_12 3clo_A 258 3 0.029 15.000 197:237 (122-358:38-234)
# OvC 45.08 M_04 z0ct 240 0 6e-37 13.000 165:224 (170-366:5-215)
PDB: 3c3w PDBsum
HEADER TRANSCRIPTION 28-JAN-08 3C3W
TITLE CRYSTAL STRUCTURE OF THE MYCOBACTERIUM TUBERCULOSIS HYPOXIC
TITLE 2 RESPONSE REGULATOR DOSR
SOURCE 2 ORGANISM_SCIENTIFIC: MYCOBACTERIUM TUBERCULOSIS;
SOURCE 3 ORGANISM_TAXID: 1773;
KEYWDS RESPONSE REGULATOR, TWO-COMPONENT REGULATORY SYSTEM, DNA-
KEYWDS 2 BINDING PROTEIN, TUBERCULOSIS, TRANSCRIPTION, TRANSCRIPTION
KEYWDS 3 REGULATION
SCOP: none
PDB: 1a04 PDBsum
HEADER SIGNAL TRANSDUCTION PROTEIN 08-DEC-97 1A04
TITLE THE STRUCTURE OF THE NITRATE/NITRITE RESPONSE REGULATOR
TITLE 2 PROTEIN NARL IN THE MONOCLINIC C2 CRYSTAL FORM
SOURCE 2 ORGANISM_SCIENTIFIC: ESCHERICHIA COLI;
SOURCE 3 ORGANISM_TAXID: 562;
KEYWDS SIGNAL TRANSDUCTION PROTEIN, RESPONSE REGULATORS, TWO-
KEYWDS 2 COMPONENT SYSTEMS
SCOP: a.4 DNA/RNA-binding 3-helical bundle
SCOP: a.4.6 C-terminal effector domain of the bipartite response regulators
SCOP: a.4.6.2 GerE-like (LuxR/UhpA family of transcriptional regulators)
SCOP: c.23 Flavodoxin-like
SCOP: c.23.1 CheY-like
SCOP: c.23.1.1 CheY-related
PDB: 1s7o PDBsum
HEADER STRUCTURAL GENOMICS, UNKNOWN FUNCTION 29-JAN-04 1S7O
TITLE CRYSTAL STRUCTURE OF PUTATIVE DNA BINDING PROTEIN SP_1288
TITLE 2 FROM STREPTOCOCCUS PYGENES
SOURCE 2 ORGANISM_SCIENTIFIC: STREPTOCOCCUS PYOGENES SEROTYPE M3;
SOURCE 3 ORGANISM_TAXID: 301448;
KEYWDS PUTATIVE DNA BINDING PROTEIN; STRUCTURAL GENOMICS, SIGNAL
KEYWDS 2 RECOGNITION PARTICLE, RNA POLYMERASE SIGMA FACTOR, X-RAT
KEYWDS 3 CRYSTALLOGRAPHY, UPF0122, STRUCTURE FUNDED BY NIH, PSI,
KEYWDS 4 PROTEIN STRUCTURE INITIATIVE, BERKELEY STRUCTURAL GENOMICS
KEYWDS 5 CENTER, BSGC, STRUCTURAL GENOMICS, UNKNOWN FUNCTION
SCOP: a.4 DNA/RNA-binding 3-helical bundle
SCOP: a.4.13 Sigma3 and sigma4 domains of RNA polymerase sigma factors
SCOP: a.4.13.3 YlxM/p13-like
PDB: 3clo PDBsum
HEADER TRANSCRIPTION REGULATOR 19-MAR-08 3CLO
TITLE CRYSTAL STRUCTURE OF PUTATIVE TRANSCRIPTIONAL REGULATOR
TITLE 2 CONTAINING A LUXR DNA BINDING DOMAIN (NP_811094.1) FROM
TITLE 3 BACTEROIDES THETAIOTAOMICRON VPI-5482 AT 2.04 A RESOLUTION
SOURCE 2 ORGANISM_SCIENTIFIC: BACTEROIDES THETAIOTAOMICRON VPI-5482;
SOURCE 3 ORGANISM_TAXID: 226186;
KEYWDS NP_811094.1, PUTATIVE TRANSCRIPTIONAL REGULATOR CONTAINING
KEYWDS 2 A LUXR DNA BINDING DOMAIN, BACTERIAL REGULATORY PROTEINS,
KEYWDS 3 LUXR FAMILY, STRUCTURAL GENOMICS, JOINT CENTER FOR
KEYWDS 4 STRUCTURAL GENOMICS, JCSG, PROTEIN STRUCTURE INITIATIVE,
KEYWDS 5 PSI-2, DNA-BINDING, TRANSCRIPTION REGULATION, TRANSCRIPTION
KEYWDS 6 REGULATOR
SCOP: none
PDB: 1ys7 PDBsum
HEADER TRANSCRIPTION REGULATOR 07-FEB-05 1YS7
TITLE CRYSTAL STRUCTURE OF THE RESPONSE REGULATOR PROTEIN PRRA
TITLE 2 COMLEXED WITH MG2+
SOURCE 2 ORGANISM_SCIENTIFIC: MYCOBACTERIUM TUBERCULOSIS;
SOURCE 3 ORGANISM_TAXID: 1773;
KEYWDS RESPONSE REGULATOR, DNA BINDING DOMAIN, PHOSPHORYLATION,
KEYWDS 2 MYCOBACTERIUM TUBERCULOSIS STRUCTURAL PROTEOMICS PROJECT,
KEYWDS 3 XMTB, STRUCTURAL GENOMICS, TRANSCRIPTION REGULATOR
SCOP: a.4 DNA/RNA-binding 3-helical bundle
SCOP: a.4.6 C-terminal effector domain of the bipartite response regulators
SCOP: a.4.6.1 PhoB-like
SCOP: c.23 Flavodoxin-like
SCOP: c.23.1 CheY-like
SCOP: c.23.1.1 CheY-related