##########
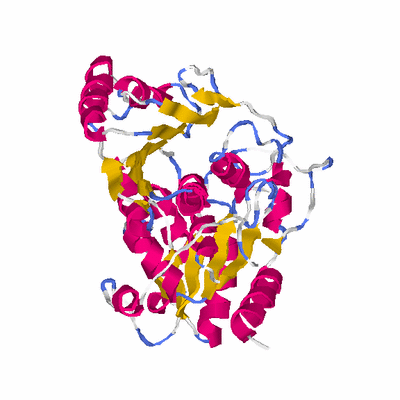
# PROTEIN: Q2246_545_343.5wLII_11276_129 344 343 # Category_B
# Length: 344
# Weight: 38612.02
# Similarity: UniProt_B6ANT2, BLAST_B6ANT2, PDB_templates_INFO
# Best Pcover Model PDB N_AA SISC E-val Seq_ID LAL Overlap
# Cat 92.44 M_00 z00p 354 0 2e-64 21.000 318:348 (3-342:2-344)
# Evl 91.86 M_00 z00p 354 0 1e-75 21.000 316:350 (3-342:2-344)
# Sid 78.78 M_01 2gt1_A 326 2 2e-10 25.000 271:300 (3-291:2-283)
# Cov 93.02 M_00 z00p 354 0 3e-65 20.000 320:348 (3-342:2-344)
# LaL 29.65 M_07 3beo_A 375 2 7e-06 18.000 102:102 (185-286:206-307)
# OvN 82.27 M_04 3dzc_A 396 2 9e-46 9.000 283:321 (1-291:25-337)
# OvC 91.86 M_00 z00p 354 0 1e-75 21.000 316:350 (3-342:2-344)
PDB: 1psw PDBsum
HEADER TRANSFERASE 21-JUN-03 1PSW
TITLE STRUCTURE OF E. COLI ADP-HEPTOSE LPS HEPTOSYLTRANSFERASE II
SOURCE 2 ORGANISM_SCIENTIFIC: ESCHERICHIA COLI;
SOURCE 3 ORGANISM_TAXID: 562;
COMPND 6 EC: 2.4.99.-;
KEYWDS STRUCTURAL GENOMICS, T832, NYSGXRC, TRANSFERASE, LPS
KEYWDS 2 BIOSYNTHETIC PATHWAY, PSI, PROTEIN STRUCTURE INITIATIVE,
KEYWDS 3 NEW YORK SGX RESEARCH CENTER FOR STRUCTURAL GENOMICS
SCOP: c.87 UDP-Glycosyltransferase/glycogen phosphorylase
SCOP: c.87.1 UDP-Glycosyltransferase/glycogen phosphorylase
SCOP: c.87.1.7 ADP-heptose LPS heptosyltransferase II
PDB: 2gt1 PDBsum
HEADER TRANSFERASE 27-APR-06 2GT1
TITLE E. COLI HEPTOSYLTRANSFERASE WAAC.
SOURCE 2 ORGANISM_SCIENTIFIC: ESCHERICHIA COLI UTI89;
SOURCE 3 ORGANISM_TAXID: 364106;
KEYWDS GT-B FOLD, TRANSFERASE
SCOP: none
PDB: 3beo PDBsum
HEADER ISOMERASE 19-NOV-07 3BEO
TITLE A STRUCTURAL BASIS FOR THE ALLOSTERIC REGULATION OF NON-
TITLE 2 HYDROLYZING UDP-GLCNAC 2-EPIMERASES
SOURCE 2 ORGANISM_SCIENTIFIC: BACILLUS ANTHRACIS;
SOURCE 3 ORGANISM_TAXID: 1392;
COMPND 4 EC: 5.1.3.14;
KEYWDS EPIMERASE, UDP-GLCNAC, ALLOSTERIC, REGULATION, ISOMERASE
SCOP: none
PDB: 3dzc PDBsum
HEADER ISOMERASE 29-JUL-08 3DZC
TITLE 2.35 ANGSTROM RESOLUTION STRUCTURE OF WECB (VC0917), A UDP-
TITLE 2 N-ACETYLGLUCOSAMINE 2-EPIMERASE FROM VIBRIO CHOLERAE.
SOURCE 2 ORGANISM_SCIENTIFIC: VIBRIO CHOLERAE;
SOURCE 3 ORGANISM_TAXID: 666;
COMPND 4 EC: 5.1.3.14;
KEYWDS UDP-N-ACETYLGLUCOSAMINE 2-EPIMERASE, STRUCTURAL GENOMICS,
KEYWDS 2 INFECTIOUS DISEASES, ISOMERASE, CENTER FOR STRUCTURAL
KEYWDS 3 GENOMICS OF INFECTIOUS DISEASES, CSGID
SCOP: none