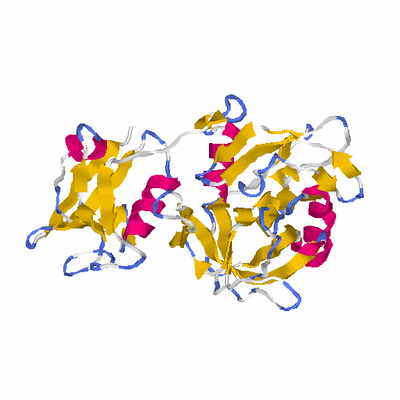##########
# PROTEIN: Q2246_545_35.5wLII_10822_10 323 35 # Category_C1
# Length: 323
# Weight: 35380.79
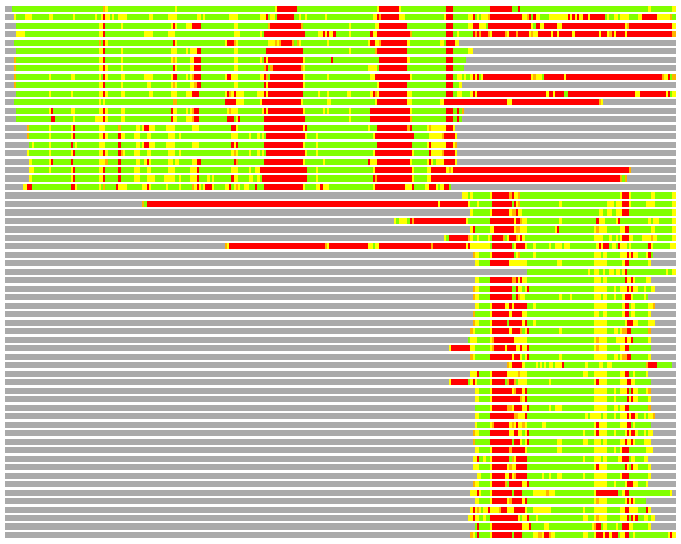
# Similarity: UniProt_A3ERG0, BLAST_A3ERG0, PDB_templates_INFO
# Best Pcover Model PDB N_AA SISC E-val Seq_ID LAL Overlap
# Cat 83.28 M_07 2z9i_A 324 6 3e-58 15.000 269:325 (17-322:4-311)
# Evl 82.04 M_01 3cs0_A 448 1 3e-88 13.000 265:288 (39-323:88-355)
# Sid 15.79 M_00 2hga_A 125 1 0.005 44.000 51:56 (251-305:21-74)
# Cov 93.50 M_04 1te0_A 318 2 8e-73 11.000 302:327 (3-323:6-313)
# LaL 27.24 M_30 2v90_A 96 6 3e-07 14.000 88:88 (233-320:9-96)
# OvN 93.50 M_04 1te0_A 318 2 8e-73 11.000 302:327 (3-323:6-313)
# OvC 82.04 M_01 3cs0_A 448 1 3e-88 13.000 265:288 (39-323:88-355)
PDB: 2z9i PDBsum
HEADER HYDROLASE 20-SEP-07 2Z9I
TITLE CRYSTAL STRUCTURE OF RV0983 FROM MYCOBACTERIUM TUBERCULOSIS-
TITLE 2 PROTEOLYTICALLY ACTIVE FORM
SOURCE 2 ORGANISM_SCIENTIFIC: MYCOBACTERIUM TUBERCULOSIS;
SOURCE 3 ORGANISM_TAXID: 1773;
SOURCE 5 ORGANISM_SCIENTIFIC: MYCOBACTERIUM TUBERCULOSIS;
COMPND 6 EC: 3.4.21.-;
KEYWDS SERINE PROTEASE, HTRA, HYDROLASE
SCOP: none
PDB: 3cs0 PDBsum
HEADER HYDROLASE 08-APR-08 3CS0
TITLE CRYSTAL STRUCTURE OF DEGP24
SOURCE 2 ORGANISM_SCIENTIFIC: ESCHERICHIA COLI;
COMPND 5 EC: 3.4.21.-;
KEYWDS DEGP, HTRA, PROTEASE, CHAPERONE, PDZ, OUTER MEMBRANE
KEYWDS 2 PROTEIN, OMP, PERIPLASM, HYDROLASE
SCOP: none
PDB: 2hga PDBsum
HEADER STRUCTURAL GENOMICS, UNKNOWN FUNCTION 26-JUN-06 2HGA
TITLE SOLUTION NMR STRUCTURE OF CONSERVED PROTEIN MTH1368,
TITLE 2 NORTHEAST STRUCTURAL GENOMICS CONSORTIUM TARGET TT821A
SOURCE 2 ORGANISM_SCIENTIFIC: METHANOTHERMOBACTER
SOURCE 4 ORGANISM_TAXID: 145262;
KEYWDS CONSERVED PROTEIN MTH1368, GFT NMR, STRUCTURAL GENOMICS,
KEYWDS 2 PSI, PROTEIN STRUCTURE INITIATIVE, NORTHEAST STRUCTURAL
KEYWDS 3 GENOMICS CONSORTIUM, NESG, HYDROLASE, METAL-BINDING,
KEYWDS 4 METALLOPROTEASE, PROTEASE, TRANSMEMBRANE, ZINC B, UNKNOWN
KEYWDS 5 FUNCTION
SCOP: none
PDB: 1te0 PDBsum
HEADER HYDROLASE 24-MAY-04 1TE0
TITLE STRUCTURAL ANALYSIS OF DEGS, A STRESS SENSOR OF THE
TITLE 2 BACTERIAL PERIPLASM
SOURCE 2 ORGANISM_SCIENTIFIC: ESCHERICHIA COLI;
SOURCE 3 ORGANISM_TAXID: 562;
COMPND 4 EC: 3.4.21.-;
KEYWDS X-RAY STRUCTURE, TWO DOMAINS, SERINE PROTEASE, PDZ, ALPHA-
KEYWDS 2 BETA PROTEIN, HYDROLASE
SCOP: b.36 PDZ domain-like
SCOP: b.36.1 PDZ domain-like
SCOP: b.36.1.4 HtrA-like serine proteases
SCOP: b.47 Trypsin-like serine proteases
SCOP: b.47.1 Trypsin-like serine proteases
SCOP: b.47.1.1 Prokaryotic proteases
PDB: 2v90 PDBsum
HEADER PROTEIN-BINDING 16-AUG-07 2V90
TITLE CRYSTAL STRUCTURE OF THE 3RD PDZ DOMAIN OF INTESTINE- AND
TITLE 2 KIDNEY-ENRICHED PDZ DOMAIN IKEPP (PDZD3)
SOURCE 2 ORGANISM_SCIENTIFIC: HOMO SAPIENS;
SOURCE 3 ORGANISM_COMMON: HUMAN;
SOURCE 4 ORGANISM_TAXID: 9606;
KEYWDS ALTERNATIVE SPLICING, PDZD3, MEMBRANE, CYTOPLASM, PDZ
KEYWDS 2 DOMAIN, PROTEIN-BINDING
SCOP: none