##########
# PROTEIN: Q2246_545_458.5wLII_11346_8 93 458 # Category_C1
# Length: 93
# Weight: 10133.71
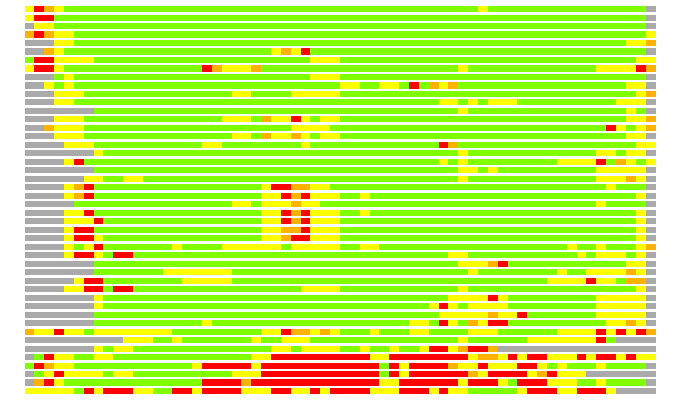
# Similarity: UniProt_A3ETU0, BLAST_A3ETU0, PDB_templates_INFO
# Best Pcover Model PDB N_AA SISC E-val Seq_ID LAL Overlap
# Cat 62.37 M_00 1y7y_A 74 2 5e-13 23.000 58:60 (31-88:9-68)
# Evl 62.37 M_00 1b0n_A 111 1 6e-15 14.000 58:61 (35-92:1-61)
# Sid 60.22 M_29 2ofy_A 86 2 8e-04 28.000 56:57 (34-89:15-71)
# Cov 86.02 M_30 1e0r_B 159 1 0.038 18.000 80:92 (3-89:33-117)
# LaL 56.99 M_10 3dnv_B 88 2 3e-08 15.000 53:53 (39-91:18-70)
# OvN 69.89 M_09 3f52_E 117 8 6e-09 19.000 65:92 (1-90:1-85)
# OvC 69.89 M_17 2ofy_A 86 2 5e-06 27.000 65:66 (29-93:10-75)
PDB: 1y7y PDBsum
HEADER TRANSCRIPTION REGULATOR 10-DEC-04 1Y7Y
TITLE HIGH-RESOLUTION CRYSTAL STRUCTURE OF THE RESTRICTION-
TITLE 2 MODIFICATION CONTROLLER PROTEIN C.AHDI FROM AEROMONAS
TITLE 3 HYDROPHILA
SOURCE 2 ORGANISM_SCIENTIFIC: AEROMONAS HYDROPHILA;
SOURCE 3 ORGANISM_TAXID: 644;
KEYWDS HELIX-TURN-HELIX, DNA-BINDING PROTEIN, TRANSCRIPTIONAL
KEYWDS 2 REGULATOR, TRANSCRIPTION REGULATOR
SCOP: a.35 lambda repressor-like DNA-binding domains
SCOP: a.35.1 lambda repressor-like DNA-binding domains
SCOP: a.35.1.3 SinR domain-like
PDB: 1b0n PDBsum
HEADER TRANSCRIPTION REGULATOR 11-NOV-98 1B0N
TITLE SINR PROTEIN/SINI PROTEIN COMPLEX
SOURCE 2 ORGANISM_SCIENTIFIC: BACILLUS SUBTILIS;
SOURCE 3 ORGANISM_TAXID: 1423;
SOURCE 11 ORGANISM_SCIENTIFIC: BACILLUS SUBTILIS;
KEYWDS TRANSCRIPTION REGULATOR, ANTAGONIST, SPORULATION
SCOP: a.34 Dimerisation interlock
SCOP: a.34.1 SinR repressor dimerisation domain-like
SCOP: a.34.1.1 SinR repressor dimerisation domain-like
SCOP: a.35 lambda repressor-like DNA-binding domains
SCOP: a.35.1 lambda repressor-like DNA-binding domains
SCOP: a.35.1.3 SinR domain-like
PDB: 2ofy PDBsum
HEADER TRANSCRIPTION 04-JAN-07 2OFY
TITLE CRYSTAL STRUCTURE OF PUTATIVE XRE-FAMILY TRANSCRIPTIONAL
TITLE 2 REGULATOR FROM RHODOCOCCUS SP.
SOURCE 2 ORGANISM_SCIENTIFIC: RHODOCOCCUS SP. RHA1;
SOURCE 3 ORGANISM_TAXID: 101510;
KEYWDS TRANSCRIPTION REGULATOR, XRE-FAMILY, STRUCTURAL GENOMICS,
KEYWDS 2 PSI, PROTEIN STRUCTURE INITIATIVE, MIDWEST CENTER FOR
KEYWDS 3 STRUCTURAL GENOMICS, MCSG
SCOP: none
PDB: 1e0r PDBsum
HEADER CHAPERONIN 06-APR-00 1E0R
TITLE BETA-APICAL DOMAIN OF THERMOSOME
SOURCE 2 ORGANISM_SCIENTIFIC: THERMOPLASMA ACIDOPHILUM;
SOURCE 3 ORGANISM_TAXID: 2303;
KEYWDS CHAPERONIN, HSP60, THERMOSOME, TCP1, GROEL, THERMOPLASMA
KEYWDS 2 ACIDOPHILUM
SCOP: c.8 The "swivelling" beta/beta/alpha domain
SCOP: c.8.5 GroEL apical domain-like
SCOP: c.8.5.2 Group II chaperonin (CCT, TRIC), apical domain
PDB: 3dnv PDBsum
HEADER TRANSCRIPTION/DNA 02-JUL-08 3DNV
TITLE MDT PROTEIN
SOURCE 2 ORGANISM_SCIENTIFIC: ESCHERICHIA COLI;
SOURCE 3 ORGANISM_TAXID: 83333;
SOURCE 11 ORGANISM_SCIENTIFIC: ESCHERICHIA COLI;
KEYWDS PERSISTENCE, MDT, DNA-BINDING, REPRESSOR, TRANSCRIPTION,
KEYWDS 2 TRANSCRIPTION REGULATION, TRANSCRIPTION/DNA COMPLEX
SCOP: none
PDB: 3f52 PDBsum
HEADER TRANSCRIPTION ACTIVATOR 03-NOV-08 3F52
TITLE CRYSTAL STRUCTURE OF THE CLP GENE REGULATOR CLGR FROM C.
TITLE 2 GLUTAMICUM
SOURCE 2 ORGANISM_SCIENTIFIC: CORYNEBACTERIUM GLUTAMICUM;
SOURCE 3 ORGANISM_COMMON: BREVIBACTERIUM FLAVUM;
SOURCE 4 ORGANISM_TAXID: 1718;
KEYWDS GENE REGULATOR, HELIX-TURN-HELIX MOTIF, TRANSCRIPTIONAL
KEYWDS 2 ACTIVATOR, HUMAN PATHOGEN, TRANSCRIPTION ACTIVATOR
SCOP: none