##########
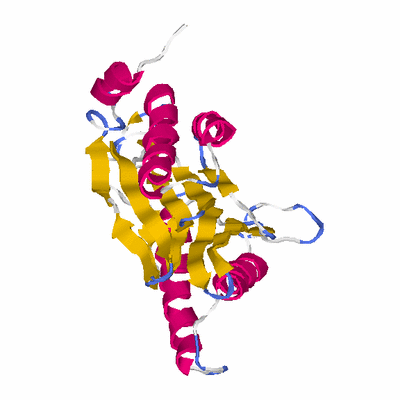
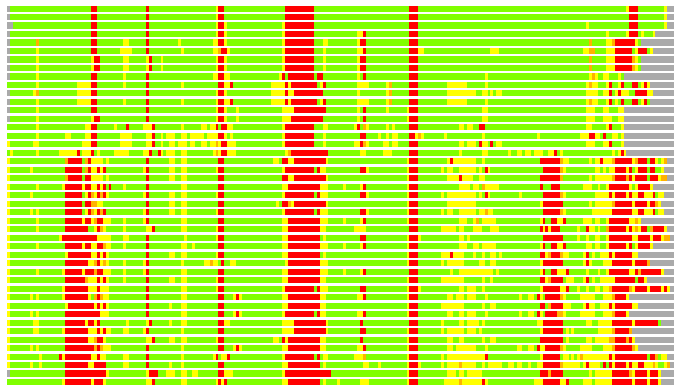
# PROTEIN: Q2246_545_547.5wLII_11391_11 251 547 # Category_C1
# Length: 251
# Weight: 27888.70
# Similarity: UniProt_B6AS34, BLAST_B6AS34, PDB_templates_INFO
# Best Pcover Model PDB N_AA SISC E-val Seq_ID LAL Overlap
# Cat 81.67 M_00 1g0u_K 212 24 2e-47 17.000 205:226 (2-227:1-205)
# Evl 78.88 M_00 1iru_L 204 2 3e-52 16.000 198:218 (2-218:1-199)
# Sid 78.49 M_07 1iru_H 205 2 1e-46 19.000 197:220 (2-220:1-198)
# Cov 87.65 M_06 1iru_I 234 2 5e-49 14.000 220:245 (2-246:1-222)
# LaL 85.66 M_30 1ryp_E 242 22 3e-37 10.000 215:226 (2-226:27-242)
# OvN 79.28 M_39 1iru_K 201 2 1e-35 12.000 199:216 (1-216:1-200)
# OvC 80.08 M_38 1iru_F 263 2 9e-38 12.000 201:254 (4-251:35-260)
PDB: 1g0u PDBsum
HEADER HYDROLASE 09-OCT-00 1G0U
TITLE A GATED CHANNEL INTO THE PROTEASOME CORE PARTICLE
SOURCE 2 ORGANISM_SCIENTIFIC: SACCHAROMYCES CEREVISIAE;
SOURCE 3 ORGANISM_COMMON: BAKER'S YEAST;
SOURCE 4 ORGANISM_TAXID: 4932;
COMPND 6 EC: 3.4.99.46;
COMPND 13 EC: 3.4.99.46;
COMPND 20 EC: 3.4.99.46;
COMPND 27 EC: 3.4.99.46;
COMPND 34 EC: 3.4.99.46;
COMPND 41 EC: 3.4.99.46;
COMPND 49 EC: 3.4.99.46;
COMPND 56 EC: 3.4.99.46;
COMPND 63 EC: 3.4.99.46;
COMPND 70 EC: 3.4.99.46;
COMPND 77 EC: 3.4.99.46;
COMPND 83 EC: 3.4.99.46;
COMPND 90 EC: 3.4.99.46;
COMPND 97 EC: 3.4.99.46;
KEYWDS PROTEASOME, UBIQUITIN, DEGRADATION, PROTEASE, NTN-HYDROLASE
SCOP: d.153 Ntn hydrolase-like
SCOP: d.153.1 N-terminal nucleophile aminohydrolases (Ntn hydrolases)
SCOP: d.153.1.4 Proteasome subunits
PDB: 1iru PDBsum
HEADER HYDROLASE 24-OCT-01 1IRU
TITLE CRYSTAL STRUCTURE OF THE MAMMALIAN 20S PROTEASOME AT 2.75 A
TITLE 2 RESOLUTION
SOURCE 2 ORGANISM_SCIENTIFIC: BOS TAURUS;
SOURCE 3 ORGANISM_COMMON: CATTLE;
SOURCE 4 ORGANISM_TAXID: 9913;
KEYWDS 20S PROTEASOME, CELL CYCLE, IMMUNE RESPONSE, PROTEOLYSIS,
KEYWDS 2 UBIQUITIN, HYDROLASE
SCOP: d.153 Ntn hydrolase-like
SCOP: d.153.1 N-terminal nucleophile aminohydrolases (Ntn hydrolases)
SCOP: d.153.1.4 Proteasome subunits
PDB: 1ryp PDBsum
HEADER MULTICATALYTIC PROTEINASE 26-FEB-97 1RYP
TITLE CRYSTAL STRUCTURE OF THE 20S PROTEASOME FROM YEAST AT 2.4
TITLE 2 ANGSTROMS RESOLUTION
SOURCE 2 ORGANISM_SCIENTIFIC: SACCHAROMYCES CEREVISIAE;
SOURCE 3 ORGANISM_COMMON: BAKER'S YEAST;
SOURCE 4 ORGANISM_TAXID: 4932;
COMPND 4 EC: 3.4.99.46;
COMPND 9 EC: 3.4.99.46;
COMPND 14 EC: 3.4.99.46;
COMPND 19 EC: 3.4.99.46;
COMPND 24 EC: 3.4.99.46;
COMPND 29 EC: 3.4.99.46;
COMPND 34 EC: 3.4.99.46;
COMPND 39 EC: 3.4.99.46;
COMPND 44 EC: 3.4.99.46;
COMPND 49 EC: 3.4.99.46;
COMPND 54 EC: 3.4.99.46;
COMPND 59 EC: 3.4.99.46;
COMPND 64 EC: 3.4.99.46;
COMPND 69 EC: 3.4.99.46;
KEYWDS MULTICATALYTIC PROTEINASE, 20S PROTEASOME, PROTEIN
KEYWDS 2 DEGRADATION, ANTIGEN PROCESSING, HYDROLASE, PROTEASE
SCOP: d.153 Ntn hydrolase-like
SCOP: d.153.1 N-terminal nucleophile aminohydrolases (Ntn hydrolases)
SCOP: d.153.1.4 Proteasome subunits