##########
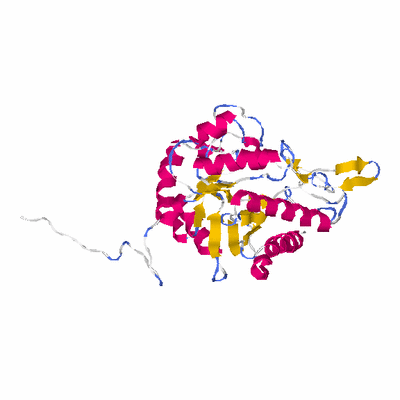
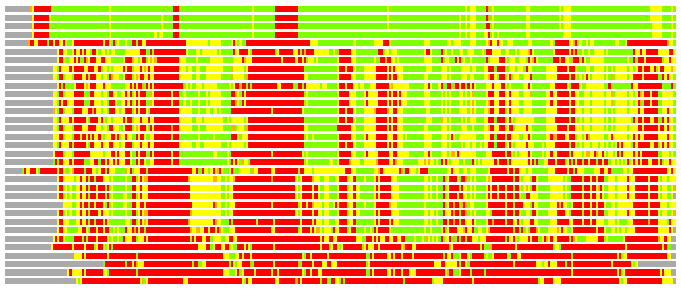
# PROTEIN: Q2246_545_55.5wLII_10954_22 650 55 # Category_C2
# Length: 650
# Weight: 73654.02
# Similarity: UniProt_B6AQE8, BLAST_B6AQE8, PDB_templates_INFO
# Best Pcover Model PDB N_AA SISC E-val Seq_ID LAL Overlap
# Cat 45.54 M_00 1uhv_A 500 6 8e-39 9.000 296:356 (225-541:2-336)
# Evl 45.54 M_00 1uhv_A 500 6 8e-39 9.000 296:356 (225-541:2-336)
# Sid 17.85 M_01 2bs9_A 503 8 1e-05 17.000 116:132 (304-429:74-195)
# Cov 45.54 M_00 1uhv_A 500 6 8e-39 9.000 296:356 (225-541:2-336)
# LaL 19.69 M_11 1egz_A 291 3 2e-04 8.000 128:132 (376-504:111-241)
# OvN 27.69 M_03 2ber_A 601 1 0.034 12.000 180:194 (87-274:319-504)
# OvC 44.31 M_14 1np2_A 436 2 0.002 10.000 288:355 (240-585:30-326)
PDB: 1uhv PDBsum
HEADER HYDROLASE 11-JUL-03 1UHV
TITLE CRYSTAL STRUCTURE OF BETA-D-XYLOSIDASE FROM
TITLE 2 THERMOANAEROBACTERIUM SACCHAROLYTICUM, A FAMILY 39
TITLE 3 GLYCOSIDE HYDROLASE
SOURCE 2 ORGANISM_SCIENTIFIC: THERMOANAEROBACTERIUM SACCHAROLYTICUM;
SOURCE 3 ORGANISM_TAXID: 28896;
COMPND 6 EC: 3.2.1.37;
KEYWDS FAMILY 39 GLYCOSIDE HYDROLASE, XYLOSIDASE, XYLAN, XYLOSE,
KEYWDS 2 COVALENT GLYCOSYL-ENZYME INTERMEDIATE
SCOP: b.71 Glycosyl hydrolase domain
SCOP: b.71.1 Glycosyl hydrolase domain
SCOP: b.71.1.2 Composite domain of glycosyl hydrolase families 5, 30, 39 and 51
SCOP: c.1 TIM beta/alpha-barrel
SCOP: c.1.8 (Trans)glycosidases
SCOP: c.1.8.3 beta-glycanases
PDB: 2bs9 PDBsum
HEADER HYDROLASE 19-MAY-05 2BS9
TITLE NATIVE CRYSTAL STRUCTURE OF A GH39 BETA-XYLOSIDASE XYNB1
TITLE 2 FROM GEOBACILLUS STEAROTHERMOPHILUS
SOURCE 2 ORGANISM_SCIENTIFIC: BACILLUS STEAROTHERMOPHILUS;
SOURCE 3 ORGANISM_TAXID: 1422;
COMPND 6 EC: 3.2.1.37;
KEYWDS HYDROLASE, BETA-XYLOSIDASE, COMPLEX OF THE COVALENT
KEYWDS 2 INTERMEDIATE, ENZYME/SUBSTRATE COMPLEX, FAMILY GH39,
KEYWDS 3 THERMOPHILIC ENZYME
SCOP: b.71 Glycosyl hydrolase domain
SCOP: b.71.1 Glycosyl hydrolase domain
SCOP: b.71.1.2 Composite domain of glycosyl hydrolase families 5, 30, 39 and 51
SCOP: c.1 TIM beta/alpha-barrel
SCOP: c.1.8 (Trans)glycosidases
SCOP: c.1.8.3 beta-glycanases
PDB: 1egz PDBsum
HEADER HYDROLASE 18-MAR-99 1EGZ
TITLE CELLULASE CEL5 FROM ERWINIA CHRYSANTHEMI, A FAMILY GH 5-2
TITLE 2 ENZYME
SOURCE 2 ORGANISM_SCIENTIFIC: ERWINIA CHRYSANTHEMI;
SOURCE 3 ORGANISM_TAXID: 556;
COMPND 6 EC: 3.2.1.4;
KEYWDS GLYCOSYL HYDROLASE; CLAN GH-A; FAMILY 5-2; CELLULASE
SCOP: c.1 TIM beta/alpha-barrel
SCOP: c.1.8 (Trans)glycosidases
SCOP: c.1.8.3 beta-glycanases
PDB: 2ber PDBsum
HEADER HYDROLASE 30-NOV-04 2BER
TITLE Y370G ACTIVE SITE MUTANT OF THE SIALIDASE FROM
TITLE 2 MICROMONOSPORA VIRIDIFACIENS IN COMPLEX WITH BETA-NEU5AC
TITLE 3 (SIALIC ACID).
SOURCE 2 ORGANISM_SCIENTIFIC: MICROMONOSPORA VIRIDIFACIENS;
SOURCE 3 ORGANISM_TAXID: 1881;
COMPND 6 EC: 3.2.1.18;
KEYWDS GLYCOSIDASE, HYDROLASE, SIALIDASE, BETA-PROPELLER,
KEYWDS 2 MICROMONOSPORA VIRIDIFACIENS
SCOP: b.1 Immunoglobulin-like beta-sandwich
SCOP: b.1.18 E set domains
SCOP: b.1.18.2 E-set domains of sugar-utilizing enzymes
SCOP: b.18 Galactose-binding domain-like
SCOP: b.18.1 Galactose-binding domain-like
SCOP: b.18.1.1 Galactose-binding domain
SCOP: b.68 6-bladed beta-propeller
SCOP: b.68.1 Sialidases
SCOP: b.68.1.1 Sialidases (neuraminidases)
PDB: 1np2 PDBsum
HEADER HYDROLASE 16-JAN-03 1NP2
TITLE CRYSTAL STRUCTURE OF THERMOSTABLE BETA-GLYCOSIDASE FROM
TITLE 2 THERMOPHILIC EUBACTERIUM THERMUS NONPROTEOLYTICUS HG102
SOURCE 2 ORGANISM_SCIENTIFIC: THERMUS NONPROTEOLYTICUS;
SOURCE 3 ORGANISM_TAXID: 116039;
COMPND 4 EC: 3.2.1.21;
KEYWDS TIM BARREL, HYDROLASE
SCOP: c.1 TIM beta/alpha-barrel
SCOP: c.1.8 (Trans)glycosidases
SCOP: c.1.8.4 Family 1 of glycosyl hydrolase