##########
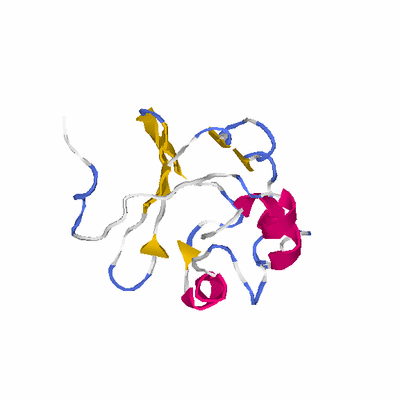
# PROTEIN: Q2246_545_69.5wLII_10961_34 184 69 # Category_C1
# Length: 184
# Weight: 19856.70
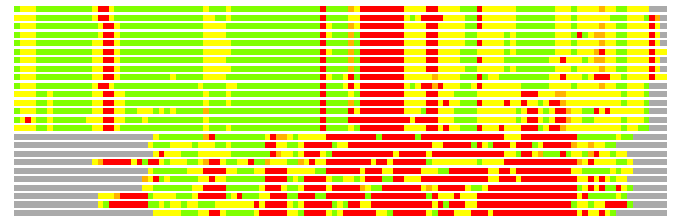
# Similarity: UniProt_B6AQ99, BLAST_B6AQ99, PDB_templates_INFO
# Best Pcover Model PDB N_AA SISC E-val Seq_ID LAL Overlap
# Cat 59.24 M_01 z0wy 438 0 5e-37 16.000 109:171 (43-153:269-437)
# Evl 61.41 M_00 z0zg 428 0 4e-37 14.000 113:161 (41-155:257-417)
# Sid 16.30 M_05 z0xs 459 0 0.063 40.000 30:30 (121-150:424-453)
# Cov 85.33 M_01 2zpd_A 190 2 5e-21 13.000 157:176 (2-166:14-181)
# LaL 26.63 M_03 z0xs 459 0 0.043 26.000 49:49 (103-151:406-454)
# OvN 83.15 M_02 z02f 202 0 5e-13 11.000 153:175 (1-164:32-200)
# OvC 68.48 M_03 2p4b_A 295 3 3e-16 9.000 126:147 (51-184:60-205)
PDB: 1q9h PDBsum
HEADER HYDROLASE 25-AUG-03 1Q9H
TITLE 3-DIMENSIONAL STRUCTURE OF NATIVE CEL7A FROM TALAROMYCES
TITLE 2 EMERSONII
SOURCE 2 ORGANISM_SCIENTIFIC: TALAROMYCES EMERSONII;
SOURCE 3 ORGANISM_TAXID: 68825;
COMPND 5 EC: 3.2.1.91
KEYWDS CELLOBIOHYDROLASE I CATALYTIC DOMAIN
SCOP: b.29 Concanavalin A-like lectins/glucanases
SCOP: b.29.1 Concanavalin A-like lectins/glucanases
SCOP: b.29.1.10 Glycosyl hydrolase family 7 catalytic core
PDB: 2a39 PDBsum
HEADER ENDOGLUCANASE 30-JAN-98 2A39
TITLE HUMICOLA INSOLENS ENDOCELLULASE EGI NATIVE STRUCTURE
SOURCE 2 ORGANISM_SCIENTIFIC: HUMICOLA INSOLENS;
SOURCE 3 ORGANISM_TAXID: 34413;
COMPND 5 EC: 3.2.1.4;
KEYWDS ENDOGLUCANASE, HYDROLASE, CELLULASE, CELLULOSE DEGRADATION,
KEYWDS 2 GLYCOSIDE HYDROLASE FAMILY 7, GLYCOSYLATED PROTEIN
SCOP: b.29 Concanavalin A-like lectins/glucanases
SCOP: b.29.1 Concanavalin A-like lectins/glucanases
SCOP: b.29.1.10 Glycosyl hydrolase family 7 catalytic core
PDB: 2zpd PDBsum
HEADER PROTEIN TRANSPORT 10-JUL-08 2ZPD
TITLE CRYSTAL STRUCTURE OF THE R43L MUTANT OF LOLA IN THE OPEN
TITLE 2 FORM
SOURCE 2 ORGANISM_SCIENTIFIC: ESCHERICHIA COLI;
SOURCE 3 ORGANISM_TAXID: 83333;
KEYWDS UNCLOSED BETA BARREL, CHAPERONE, PERIPLASM, PROTEIN
KEYWDS 2 TRANSPORT, TRANSPORT
SCOP: none
PDB: 1iwl PDBsum
HEADER PROTEIN TRANSPORT 17-MAY-02 1IWL
TITLE CRYSTAL STRUCTURE OF THE LIPOPROTEIN LOCALIZATION FACTOR,
TITLE 2 LOLA
SOURCE 2 ORGANISM_SCIENTIFIC: ESCHERICHIA COLI;
SOURCE 3 ORGANISM_TAXID: 562;
KEYWDS UNCLOSED BETA BARREL, PROTEIN TRANSPORT
SCOP: b.125 Prokaryotic lipoproteins and lipoprotein localization factors
SCOP: b.125.1 Prokaryotic lipoproteins and lipoprotein localization factors
SCOP: b.125.1.1 Outer-membrane lipoproteins carrier protein LolA
PDB: 2p4b PDBsum
HEADER SIGNALING PROTEIN 12-MAR-07 2P4B
TITLE CRYSTAL STRUCTURE OF E.COLI RSEB
SOURCE 2 ORGANISM_SCIENTIFIC: ESCHERICHIA COLI K12;
SOURCE 3 ORGANISM_TAXID: 83333;
KEYWDS OPEN AND CLOSED FORM, SIGNALING PROTEIN
SCOP: none